Erik Amézquita, PhD student in the department of Computational Mathematics, Science and Engineering at Michigan State University, is the lead author for a paper published in in silico Plants that presents a new technique to analyze shapes in plants.
Topological Data Analysis (TDA) is an emerging mathematical discipline that arises from the notion that all data is shape, and all shape is data. With TDA, the shape of diverse objects can be characterized using abstract mathematical representations based on algebraic topology. TDA approach does not depend on the existence of homologous landmarks –similar features due to shared ancestry from a common ancestor–, nor is restricted to objects of a particular orientation or dimension. TDA offers thus a robust, comprehensive, comparable, and quantifiable framework to measure shape for a diversity of inputs. One of the many TDA tools available is the Euler Characteristic Transform (ECT), which measures how the intrinsic topology –specifically the Euler characteristic- of an object changes as it is sliced across all possible thresholds and directions.
The choice of using the ECT arises from two important reasons. The first one is that the ECT computation for a small seed takes just a couple of seconds, which is important when dealing with a large volume of data. The second reason is that slicing a seed across all possible directions mathematically guarantees to encode all there is to know about shape, enough to even reconstruct the original shape from scratch.
“The caveat is that there is actually an infinite number of directions to compute. Nonetheless, even by taking 150 or so directions, we seem to encode enough morphological information to then produce exciting results,” explains Erik Amézquita, a mathematician by training now turned into a plant biologist.
The authors compared the effectiveness of using traditional shape descriptors, topological shape descriptors, or a combination of both to characterize and identify seeds of different barley accessions.

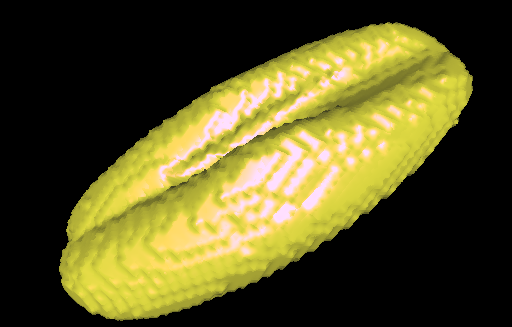
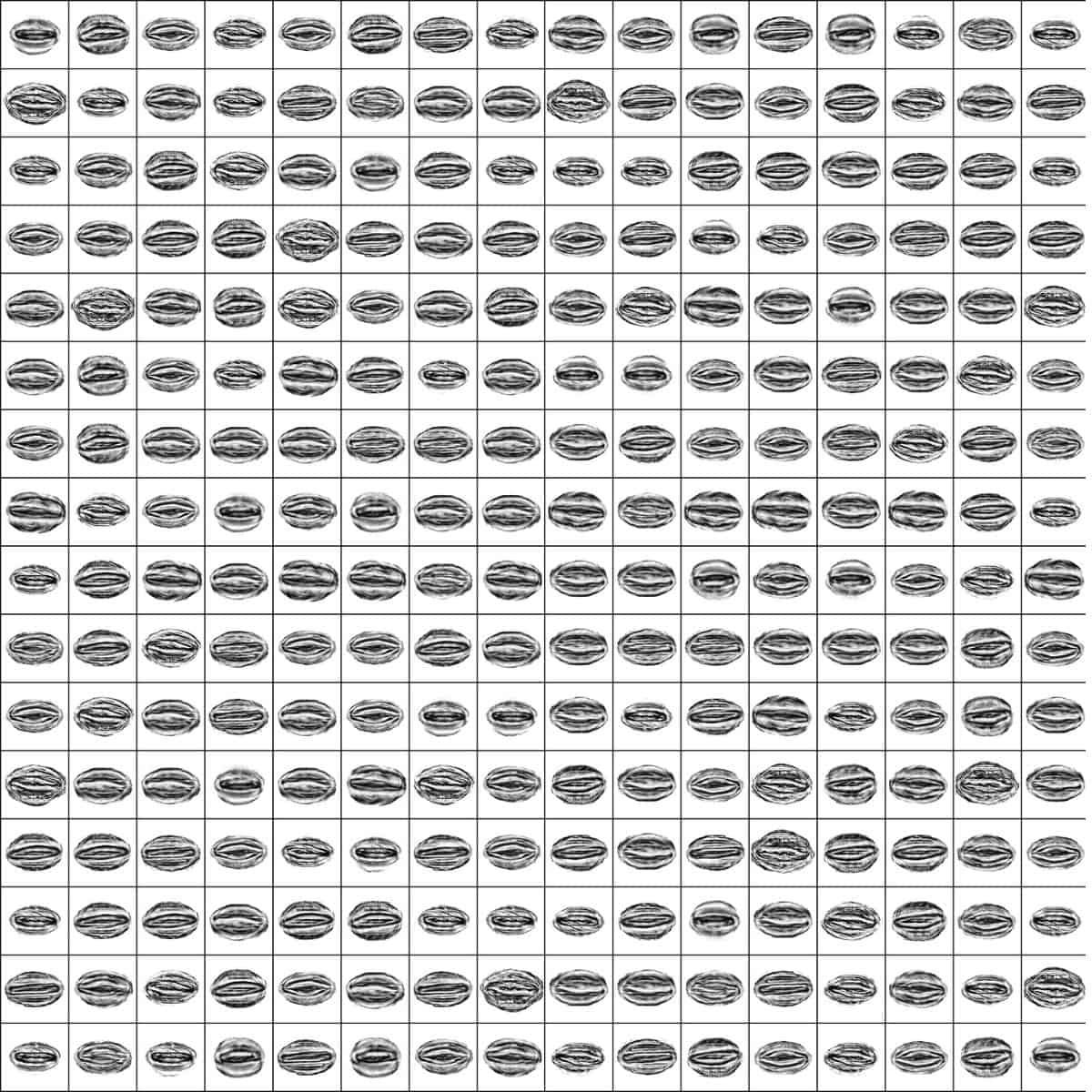
First, they collected panicles from 28 accessions with diverse spike morphologies and geographical origins. Then they scanned these panicles (seed clusters), in batches of three or four panicles each, using X-ray computed tomography (CT) (Fig. 1). These scans were later digitally processed to isolate over three thousand individual barley seeds from the panicles. Finally, every seed was aligned and oriented according to their three main principal components.
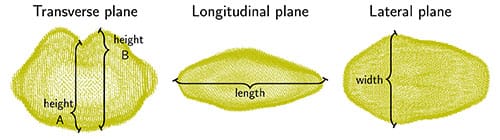
The authors proceeded then to quantify the shape of the barley grains. They first measured 11 different traditional shape descriptors, like length, width, heights, surface area, and volume for each seed (Fig. 2).
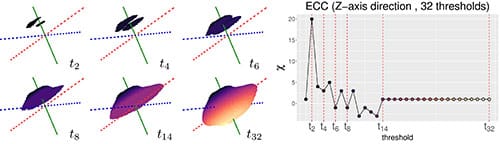
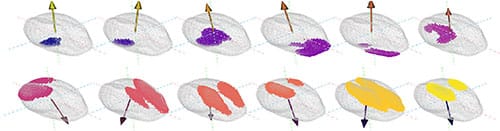
Next, the topological shape descriptors were measured with the ECT. To compute the ECT, first the seeds were chopped across a fixed direction into 16 slices of equal thickness. Next, the seeds were reconstructed by adding one slice at a time and changes in the Euler Characteristic were recorded (Fig. 3). This chopping, slice by slice reconstruction, and Euler characteristic tracking was performed for 158 different directions in total. The ECT procedure produced more than 2500 different slices, corresponding to more than 2500 topological shape descriptors for each seed. To prevent distortions caused by working with data in high dimensions – known as the curse of dimensionality, – a dimensionality reduction was necessary.
To study the suitability of all the shape descriptors, a computer was tasked with characterizing and predicting the 28 barley seed accessions solely using grain morphology information. The computer, a support vector machine, used three kinds of training. First, the machine exclusively used traditional shape descriptors. Next, the machine was trained solely with topological shape descriptors. Finally, the machine used both sources of information.
The authors found that for most of the accessions, topological features help the computer produce better prediction rates than traditional shape features. These classification results were boosted further when both traditional and topological information was combined, demonstrating that topology measures features missed by the traditional setting. Moreover, while traditional shape descriptors are able to cluster the seeds based on their accession, topological shape descriptors were able to cluster them further based on their panicle.
To determine exactly what is that “something” missed by the traditional features, several evaluations of variance analyses were undertaken. An exploration of the directions and slices used to compute the ECT reveal that the shape of the crease and bottom of the seeds discriminate accessions the most (Fig. 4)
Says Amézquita, “The Euler characteristic is a simple yet powerful way to reveal features not readily visible to the naked eye. There is hidden morphological information that traditional and geometric morphometric methods are missing. The Euler characteristic, and TDA in general, can be readily computed from any image data. TDA suggests a new exciting path, driven by morphological information alone, to further explore the phenotype-genotype relationship.”
READ THE ARTICLE:
Erik J Amézquita, Michelle Y Quigley, Tim Ophelders, Jacob B Landis, Daniel Koenig, Elizabeth Munch, Daniel H Chitwood, Measuring hidden phenotype: Quantifying the shape of barley seeds using the Euler Characteristic Transform, in silico Plants, 2021;, diab033, https://doi.org/10.1093/insilicoplants/diab033
This manuscript is part of in silico Plant’s Functional Structural Plant Model special issue.
All of the data and code used in this article are freely and openly available at https://doi.org/10.5061/dryad.rxwdbrv93 and https://github.com/amezqui3/demeter/.




