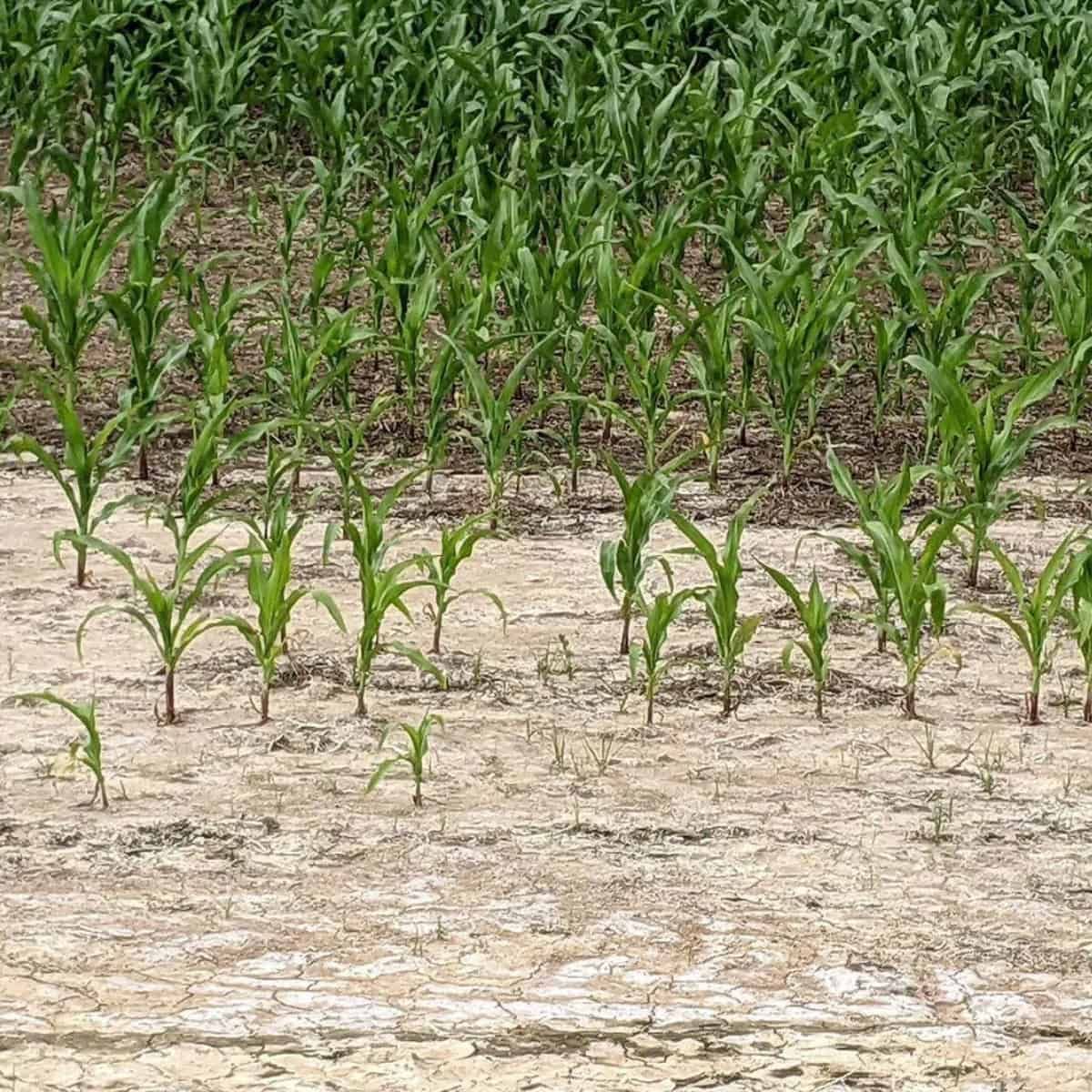
During the last decade, research activity on plant phenotyping increased exponentially. This can be largely attributed to the emergence of the new phenotyping platforms, the appearance of more complex technologies, and the increasing availability of powerful sensors.
It is believed that combining information extracted from phenotyping data with mathematical and computational models will help scientists address the urgent need to improve plant traits to ensure food security in the coming decades.
In a new study published in in silico Plants, Clément Saint Cast, then a Postdoctoral Researcher at UCLouvain, and colleagues explored how to connect phenomics and modelling communities. First, they quantified to what extent phenomics and modelling communities exchange datasets and use them together. They then suggested how this could be facilitated.
Determining the established and potential connection between communities
After identifying over 4,000 scientific papers related to plant phenotyping, plant image analysis software tools, process-based crop models, and individual plant-based models, the authors assigned papers to research topics depending on their content. Terms occurring in titles, abstracts and keywords were extracted from papers and analyzed to identify the frequency distribution of the key terms associated with the papers.
Using the VOSviewer software, they were able to visualize connections between papers from both phenomics and modelling communities. The software calculates similarities among papers based on the number of cited references they have in common. The larger the number of cited references papers have in common, the stronger the papers are related to each other.
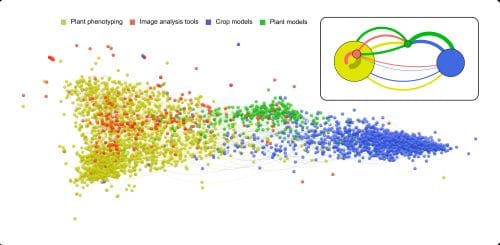
Despite the overlap in topics addressed by model and phenotyping communities, the number and proportion of cited references in common between communities are low, highlighting a low level of connection between these communities.
According to Clément Saint Cast, “this result is likely due to the different scientific goals of these communities, the lack of awareness of the benefits promoted by each community, the heterogeneous terminology used by the communities and the lack of common platforms to enable transparent data exchange.”
The authors present a 3-part strategy to move towards better connection and collaboration between phenotyping and modelling communities.
1. Raise awareness of the diversity of models, phenomics datasets and phenotyping platforms that exist.
The “Quantitative Plant” online portal (quantitative-plant.org) allows users to explore over 100 diverse plant and crop simulation models and their applications. It helps researchers search for image analysis software for their phenotyping experiments to find out potential game-changing model applications on the associated crop or plant model webpage.
The IPPN (https://www.plant-phenotyping.org/infrastructure_map) and EMPHASIS (https://emphasis.plant-phenotyping.eu/emphasis_infrastructure_map) have databases of phenotyping platforms. They provide information on existing and upcoming phenotyping platforms as well as their characteristics and uses.
2. Standardize terminology used by the communities.
Clément Saint Cast explains, “currently, the terminology used by phenotyping and modelling communities can be very heterogeneous depending on the research discipline, scale, objectives, and even between research groups. This limits the ability to accurately relate information within and across communities. We suggest a solution to facilitate the connection and the exchange of information: the use of a controlled and standardized dictionary of common and internationally recognized terms that can be shared among the communities.”
While some work has been done to tackle these issues by the phenotyping community, additional work by the plant and crop modelling communities needs to be undertaken.
3. Develop common platforms to enable transparent data exchange.
Long-term cooperation between the phenomics and modelling communities will require the development of common platforms designed to enable transparent data exchange from models to experiments and vice-versa. The development of a common hosting platform should consider (i) phenomics data with their associated metadata, (ii) phenomics data in a harmonized format and (iii) models with their associated translators and connectors to allow the connection between phenomics data and other models.
“We have undertaken this work because we believe that communication and collaboration between communities and open-source data and models provide a great opportunity for rapid advancement in plant biology.”
READ THE ARTICLE:
Clément Saint Cast, Guillaume Lobet, Llorenç Cabrera-Bosquet, Valentin Couvreur, Christophe Pradal, François Tardieu, Xavier Draye, Connecting plant phenotyping and modelling communities: lessons from science mapping and operational perspectives, in silico Plants, Volume 4, Issue 1, 2022, diac005, https://doi.org/10.1093/insilicoplants/diac005
You can read more about the Quantitative Plant website on Botany One.



