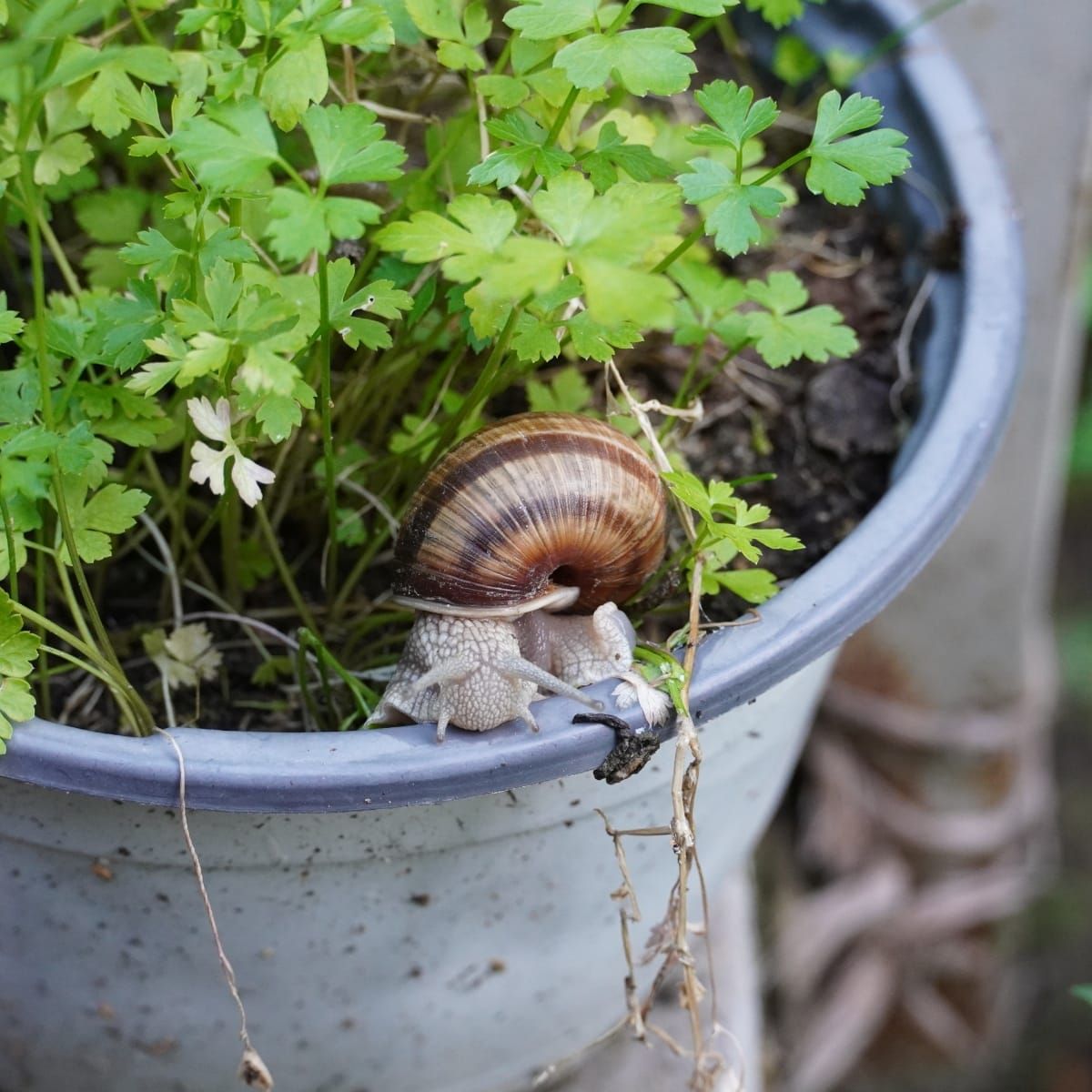
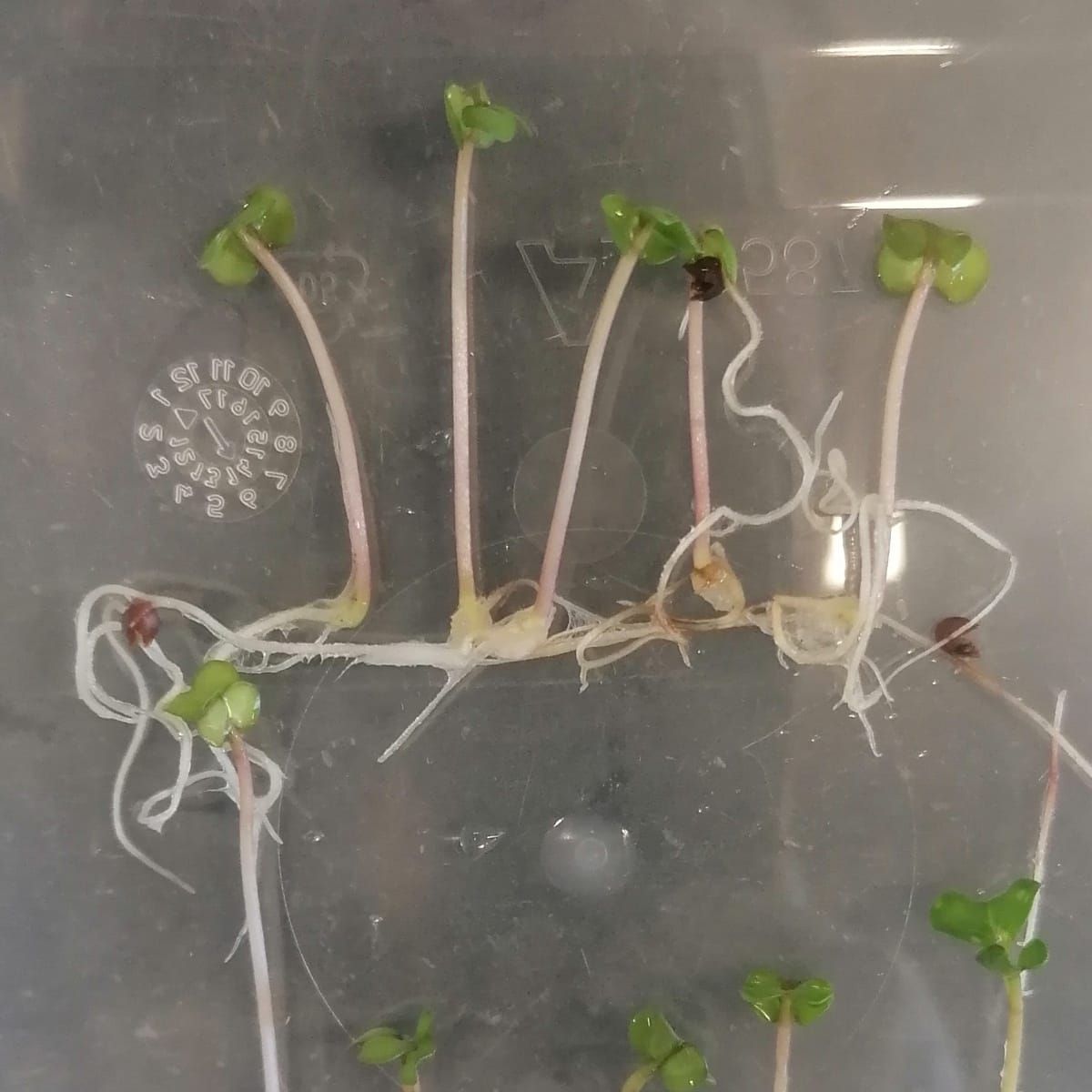
Increasing crop production is at the core of achieving food security in the 21st century. A group of scientists is addressing this challenge by studying shoot branching, one of the key factors affecting yield. They simulated breeding trials with selection for branching using computational simulations ofgene-controlled traits. They found that selection for branching is difficult due to the complex interactions among the traits hormones and sucrose that control branching.
Despite the detailed knowledge of the molecular and physiological mechanisms of branching, translating these discoveries into breeding outcomes is still a challenge. Examining the gene to phenotype network of complex traits using prior knowledge of biological interactions can unmask hidden genetic variation and identify intermediate traits that can increase the selection accuracy and efficiency of breeding for branching.
Dr. Owen Powell, Postdoctoral Research Fellow at Queensland Alliance for Agriculture and Food Innovation at the University of Queensland and coauthors developed a gene-to-phenotype network for shoot branching. Using this model, they investigated the magnitude of hidden genetic variation that control the complex trait of shoot branching and its implications for genetic gain over cycles of selection.
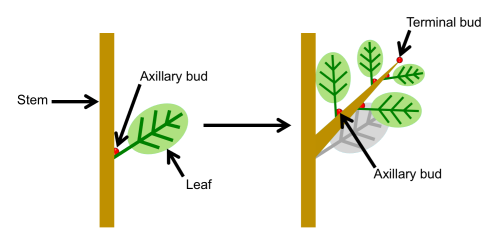
Shoot branching depends on the outgrowth of axillary buds that form in the axils of leaves. Previous research demonstrated that time to axillary bud outgrowth was controlled by the levels of hormones (auxin, cytokinins, strigolactones), and sucrose using an empirical shoot branching network model. The authors extended this model to include explicit causal genetic effects in the control of hormone and sucrose levels.
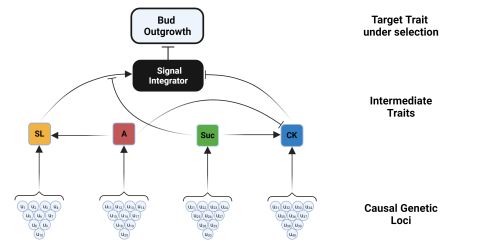
The authors performed in silico selection experiments to determine if the network view of shoot branching in plants had implications on the prediction of response to breeding selection. It did and there were plenty of surprises! Selection for the shoot branching gene to phenotype network for faster bud outgrowth was unsuccessful. Powell explains, “we expected to see diminishing responses to selection over time, eventually reaching a permanent plateau. Instead, we saw a combination of temporary and permanent plateaus, especially for sucrose. This is an indication of genetic canalization, an emergent property of the complex interactions among the components of the shoot branching gene-to-phenotype network.”
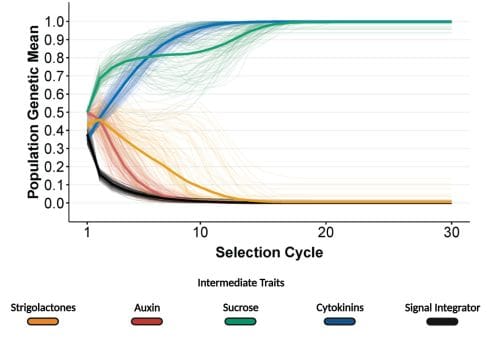
Genetic canalization describes a phenomenon of the gene-to-phenotype map when the phenotype is resistant to genetic changes, in this case caused by selection pressure.
To examine the drivers of the unexpected responses to selection, they then examined the patterns of hormone and sucrose levels and allele frequency of the populations over the selection cycles. “The unexpressed genetic expression for sucrose can be explained by the complex interaction between sucrose and strigolactone signalling in the gene-to-phenotype network. Multiple genetic combinations of sucrose and strigolactone produce similar values for the bud outgrowth, resulting in genotypes with completely different combinations of strigolactone and sucrose levels producing similar values for time to bud outgrowth,” Powell continued.
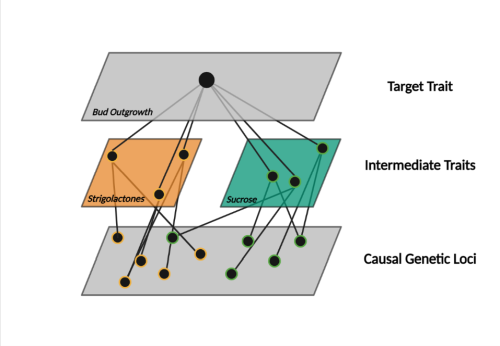
Genetic canalization is expected to be pervasive in other complex traits, such as grain yield, that result from interactions among multiple genes, traits, environments, and agronomic management practices. This illustrates the need for predictive breeding to take a network view of complex traits to advance our understanding of selection response and the efficiency of developing resilient crops for future climates.
READ THE ARTICLE:
Owen M Powell, Francois Barbier, Kai P Voss-Fels, Christine Beveridge, Mark Cooper, Investigations into the emergent properties of gene-to-phenotype networks across cycles of selection: A case study of shoot branching in plants, in silico Plants, 2022;, diac006, https://doi.org/10.1093/insilicoplants/diac006





