Crop improvement to increase yields and mitigate the effects of climate change is limited by our knowledge of how genotypes affect whole-plant phenotypes. The key will be linking genotype to phenotypes, not in abstract, statistical terms but via measurable, molecular pathways. This process is challenging, but new computational models can help us meet this challenge and provide new insights.
Dr Yin Hoon Chew, currently a Postdoctoral Research Associate at University of Birmingham, and colleagues extended an existing model to predict organism-level biomass growth from genotype based on molecular pathways in a new paper published in in silico Plants. The authors used diverse metabolic and physiological data to combine and extend mathematical models of rhythmic gene expression, photoperiod-dependence, growth, and starch metabolism within a Framework Model for the vegetative growth of Arabidopsis thaliana.
The original Arabidopsis Framework Model version 1 (FMv1) was designed to study circadian effects on physiology. It represents the interacting, physiological components of vegetative growth in Arabidopsis thaliana up to flowering, in a simple, modular fashion. The authors extended this model to include:
- a light signaling submodel,
- starch mobilization controlled by the circadian clock submodel (rather than a fixed rate),
- levels of malate and fumarate pools in the carbon submodel, and
- additional RNAs, proteins, and protein complexes within the circadian clockwork and in its links to photoperiodism.
According to Chew, “The first Arabidopsis Framework Model showed how the circadian clock affected the number of leaves by controlling flowering time, without any direct effect on the size or mass of the plant. In fact, the clock controls many processes related to plant metabolism. We expected that mis-timing those daily processes would not just affect developmental timing but also the plant’s biomass. It wasn’t clear which of the clock-regulated processes would have most effect on biomass, and that’s a question that modelling can address.”
The new Arabidopsis Framework Model version 2 (FMv2) updated the simulated partitioning of carbon from daytime photosynthesis into transient storage as starch in leaf cells, and the use of those stores to provide sugars at night. Both model versions vary the proportion of carbon that is stored, depending on the photoperiod. The ‘growth-first’ process in the FMv1 didn’t match more recent evidence, so the FMv2 has a more cautious, ‘store-first’ strategy that fits with the plants’ end-of-day starch levels across photoperiods. Rhythmic output from the clock gene circuit limits how fast the starch stores can be used, to avoid starvation before photosynthesis resumes at dawn. Introducing this control in FMv2 matched the end-of-night starch levels.
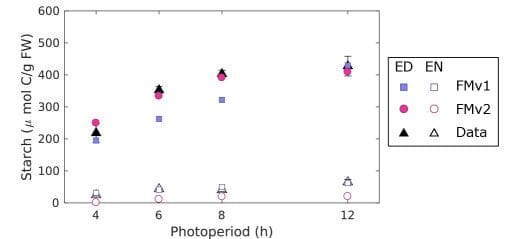
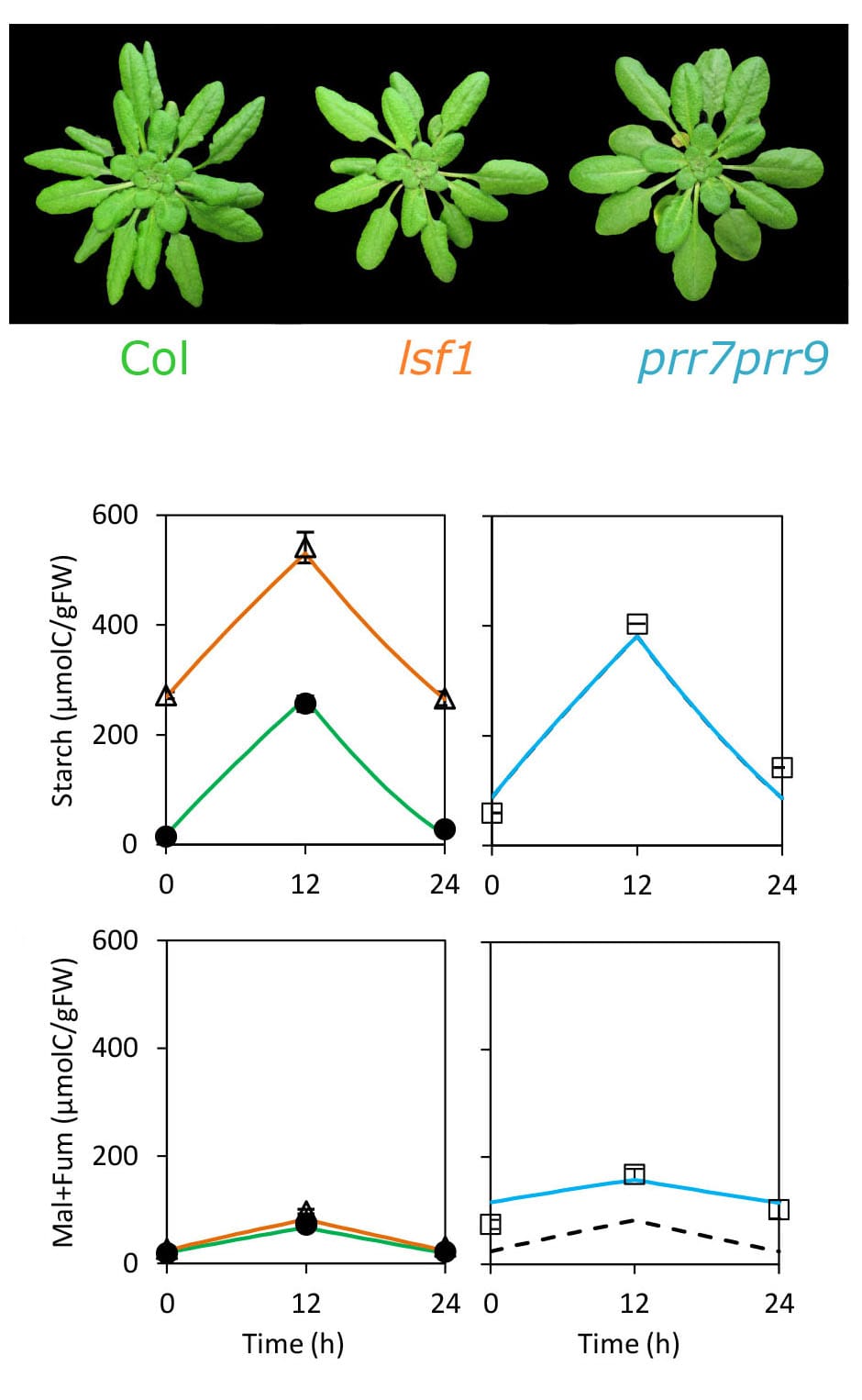
FMv2 was then tested to see if it could explain the biomass decline observed in the prr7prr9 long period mutant, which has a 28h clock. The model correctly predicted the higher starch levels observed in the prr7prr9 mutants due to their delayed circadian timing. Simulating the whole-plant context of the Framework Model allowed the authors to test whether this metabolic change was enough to reduce the biomass of the mutants compared to wild-type plants.
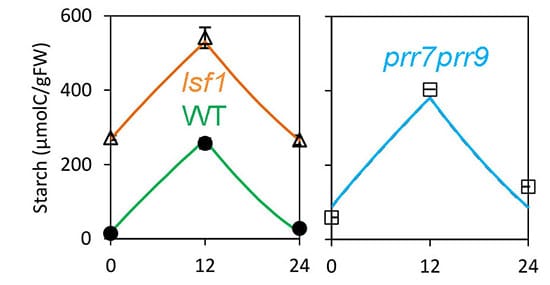
Previous work indicated that the clock affected biomass by changing the use of stored starch at night. A mild starch-use mutant, lsf1, demonstrated reduced plant biomass in line with the FMv2 prediction. The FMv2 results showed that mis-timed starch use could account for most of the biomass decline in the clock mutants too, but suggested that another clock-regulated process was also involved. The authors’ metabolite data showed that mobilization of malate and fumarate was mis-regulated in the clock mutants. Modelling both starch and these, secondary stores accounted for all the biomass defect. Interestingly, no defects in chlorophyll level and photosynthetic rates were observed in prr7prr9 plants, though they had previously been suggested to affect the growth of other clock mutants in Arabidopsis.
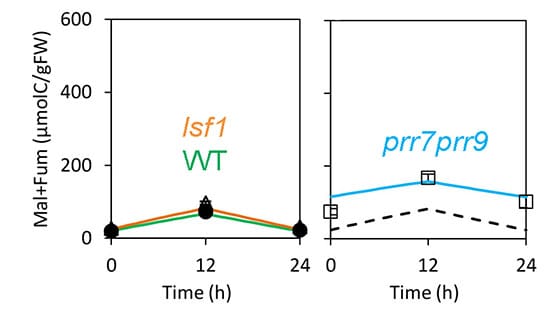
Chew concludes, “The success of the FMv2 shows that we can understand subtle effects at the whole-plant level, in this case just from changing the timing of gene expression. By ‘understand’ we mean ‘explain and predict’. We welcome Arabidopsis researchers to add to the Open Data that we’ve shared and extend this model to study their areas of expertise. Not all details of this model will transfer to crop species, but it extends the ‘proofs of principle’ for informing crop improvement at the molecular level.”
Photo of actual Wildtype, prr7prr9 and lsf1 Arabidopsis mutants, simulations of the biomass of these plants, and simulated plant biomass.
READ THE ARTICLE:
Yin Hoon Chew, Daniel D Seaton, Virginie Mengin, Anna Flis, Sam T Mugford, Gavin M George, Michael Moulin, Alastair Hume, Samuel C Zeeman, Teresa B Fitzpatrick, Alison M Smith, Mark Stitt, Andrew J Millar, The Arabidopsis Framework Model version 2 predicts the organism-level effects of circadian clock gene mis-regulation, in silico Plants, Volume 4, Issue 2, 2022, diac010, https://doi.org/10.1093/insilicoplants/diac010
This article is part of the in silico Plant Integrative and Multiscale Modelling special issue.
The Framework Model version 2 is openly available at: https://fairdomhub.org/models/248. The data used to build and test the model are shared at: https://fairdomhub.org/investigations/123. In addition, a simpler online version of FMv2 for non-experts is available at: http://turnip.bio.ed.ac.uk/fm/