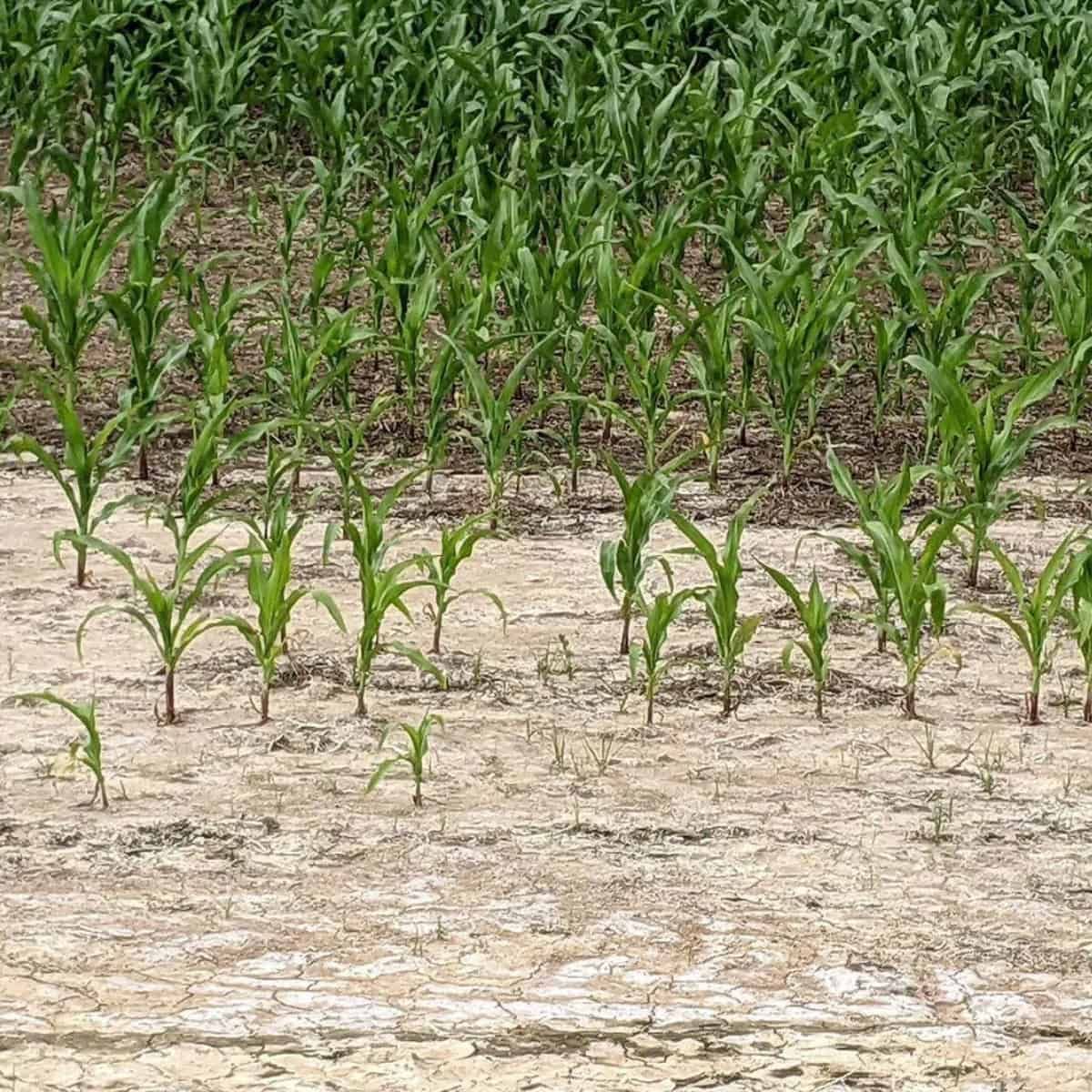
To model how plants grow and develop according to their genotypes, phenotypes, environment and/or management the use of a realistic three-dimensional (3D) canopy helps refine the predictions of light interception and photosynthesis.
The development of 3D imaging techniques for estimating canopy structure, shoot growth and biomass has expanded during the last couple of years. However, securing sufficient 3D-scanned data is time-consuming because of the need to manually process the data. In large-scale simulations, single-plant or few-plant reconstructions are often duplicated for convenience, and therefore lack phenotypic diversity. Now deep generative models can be used to learn and create realistic 3D data.
Dr. Jung Eek Son, professor of plant science at Seoul National University, and colleagues generated leaf models and extracted their traits by using deep generative models. The authors scanned pepper plants at various stages of development using a high-resolution portable 3D scanner. Point clouds were obtained from the scans, then these were used to train the deep generative models that could generate leaves.
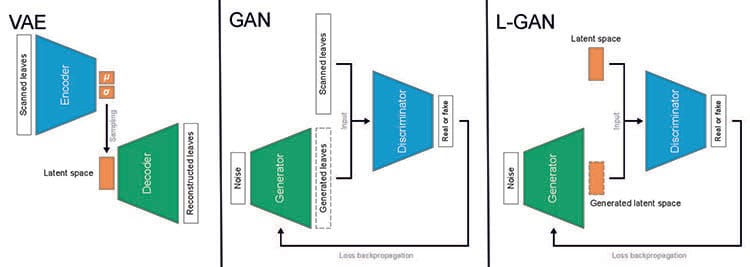
The authors compared leaves generated using three deep generative models: variational autoencoder (VAE), generative adversarial network (GAN) and latent space GAN.
While a VAE takes raw data from an image, encodes it with lower resolution, and then reconstructs it, a GAN generates an image from noise, then discriminates the image based on the raw data to determine if it is real or fake. A latent space GAN (L-GAN) has the same basic structure and training method as a GAN but uses latent variables: features from which the learning system could detect or classify patterns in the input, instead of raw data.
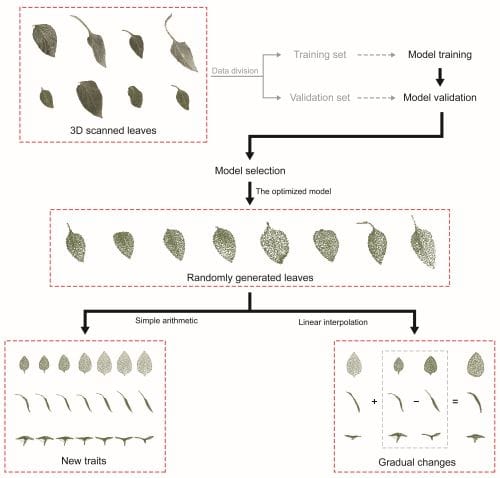
Reliable 3D phenotypes of pepper leaves were created by the deep generative models. Among the deep generative models, L- GAN showed the highest performance in generating realistic leaves. “We compared several generative models for leaf generation. In this way, the shape of leaves could be controlled with linear interpolation and simple arithmetic operations. That is, the generative model includes morphological traits somewhere in the model parameters. The first step toward the practical use of deep generative models was achieved for autonomous creations of 3D plant models without complicated feature extraction” says Son.
While the 3D shape of leaves viewed from the top were used to train the deep generative models, the models were also able to generate images to assess latent variables such as inclination and curvature from the data.
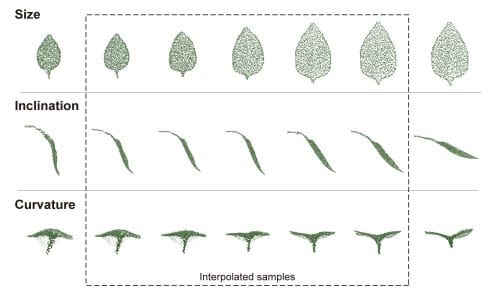
Son concludes, “deep generative models can parameterize and generate morphological traits in digitized 3D plant models and add realism and diversity to plant phenotyping studies and models.”
READ THE ARTICLE:
Taewon Moon, Hayoung Choi, Dongpil Kim, Inha Hwang, Jaewoo Kim, Jiyong Shin, Jung Eek Son, Autonomous construction of parameterizable 3D leaf models from scanned sweet pepper leaves with deep generative networks, in silico Plants, Volume 4, Issue 2, 2022, diac015, https://doi.org/10.1093/insilicoplants/diac015



