Although genome-wide association studies (GWAS) enable the identification of single nucleotide polymorphism (SNP) variants associated with traits of interest, many of the identified variants are in non-coding regions, and presumably only influence gene expression regulation. Therefore, identifying causative genes underlying a given phenotype using variants alone is very challenging. Integrating GWAS and gene coexpression networks can help prioritize high-confidence candidate genes, as the expression profiles of trait-associated genes can be used to mine novel candidates. A new article published in in silico Plants presents a statistical framework that automates the integration of gene coexpression networks and GWAS-derived SNPs to prioritize candidate genes associated with traits of interest.
Graduate Researcher Fabricio Almeida-Silva and Associate Professor Dr. Thiago Venancio at the State University of Northern Rio de Janeiro present cageminer (candidate gene miner), an R/Bioconductor package to prioritize candidate genes through the integration of GWAS and coexpression networks.
cageminer uses a guide gene-based approach to discover novel candidates that are coexpressed with known trait-associated genes and that are significantly induced or repressed in conditions of interest.
While an existing computational approach, Camoco, exists that can integrate loci identified by GWAS with functional information derived from gene coexpression networks, cageminer is able to discover candidate genes within a larger sliding window, allowing it to detect more candidate genes.
Input data required by cageminer are SNP positions, guide genes, and a gene coexpression network.
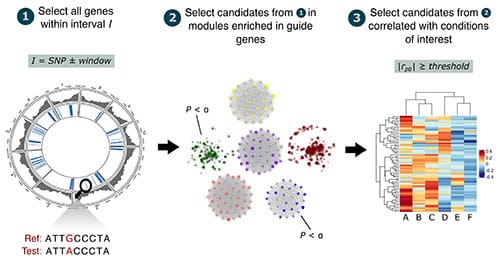
Genes are considered high-confidence candidates if they pass all three filtering criteria implemented in cageminer:
- physical proximity to SNPs,
- coexpression with known trait-associated genes, and
- significant changes in expression levels in conditions of interest.
Prioritized candidates can also be scored and ranked to select targets for experimental validation.
The authors applied cageminer to a real data set of Capsicum annuum response to Phytophthora infection to identify prioritized candidates that encode proteins related to known plant immunity-related processes. Easy-to-follow documentation of the session including code, explanations and figures are included in a supplementary file to the article.
Venancio concludes, “we developed cageminer to prioritize gene candidates, leading to a significant reduction in the size of candidate gene lists. We anticipate that this package will contribute to the advancement of population genomics and to the identification of genes for biotechnological applications.”
READ THE ARTICLE:
Fabricio Almeida-Silva, Thiago M Venancio, cageminer: an R/Bioconductor package to prioritize candidate genes by integrating GWAS and gene coexpression networks, in silico Plants, 2022; diac018, https://doi.org/10.1093/insilicoplants/diac018
All data and code used in this manuscript are freely available in a GitHub repository (https://github.com/almeidasilvaf/cageminer_benchmark) to ensure full reproducibility





