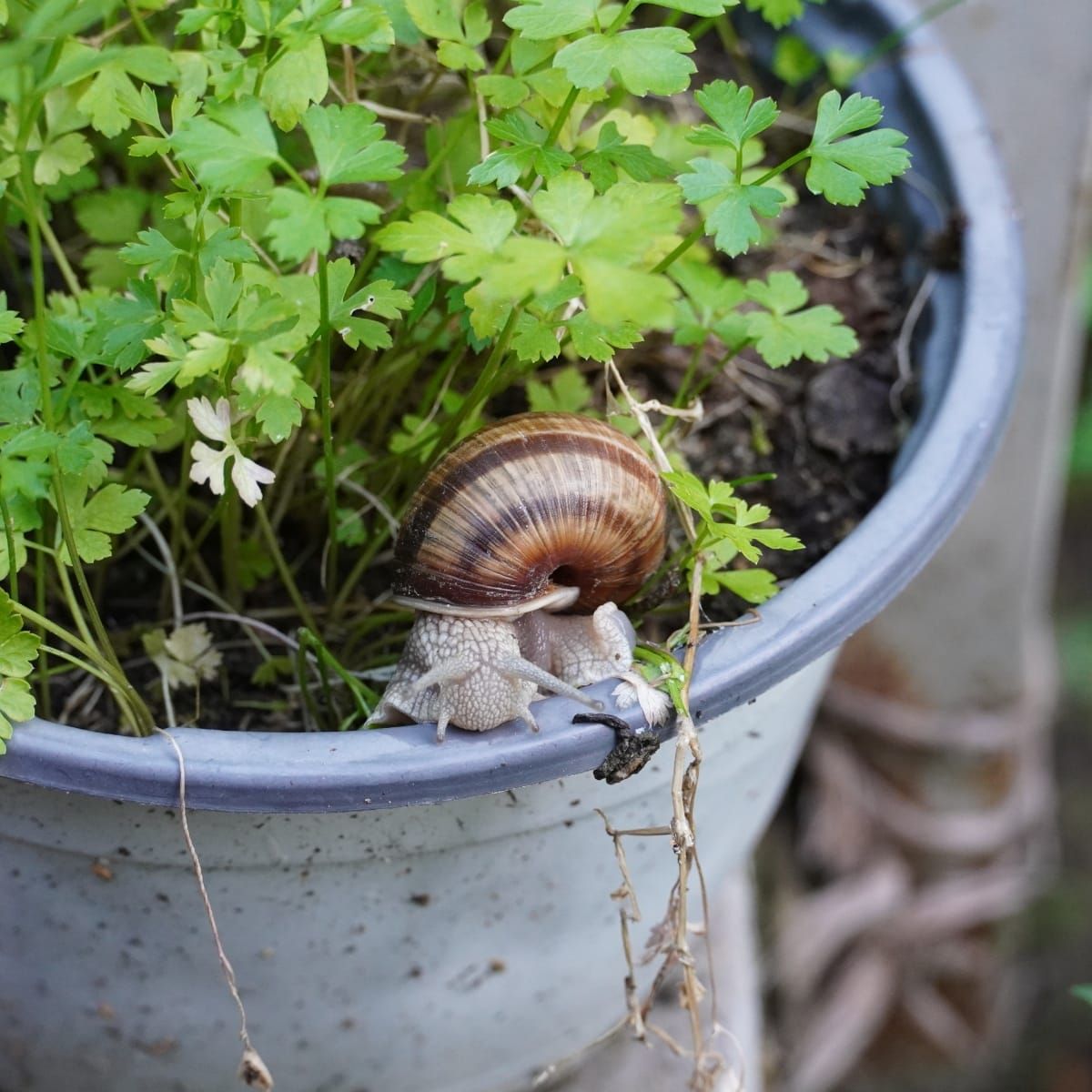
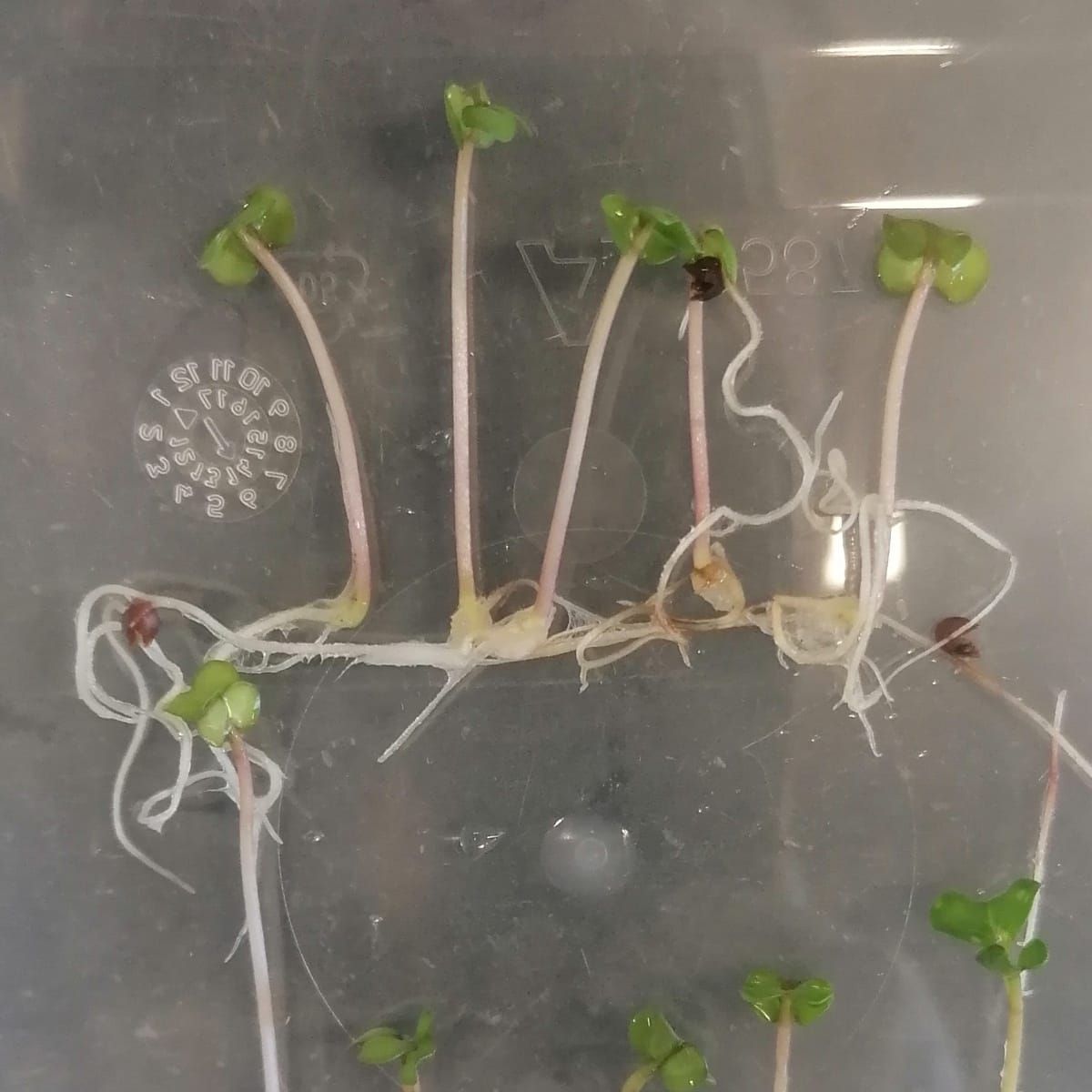
The Calvin cycle is the stage of photosynthesis that uses the energy stored by the light-dependent reactions to form glucose and other carbohydrate molecules.
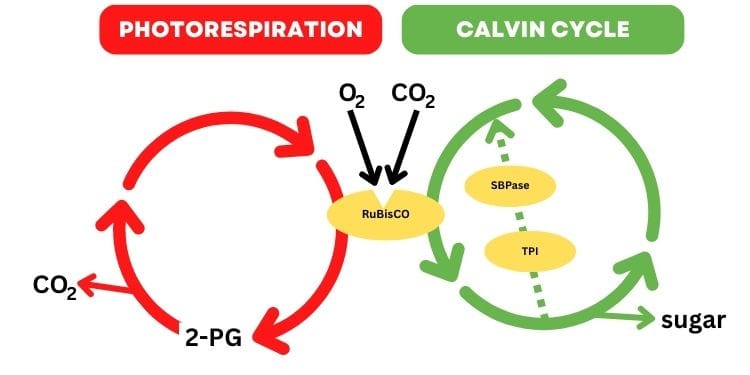
Photorespiration emerges from the competition of O2 and CO2 at the active site of the enzyme Rubisco. Oxygenation reduces rates of carboxylation, and therefore reduces the net carbon assimilation and energy efficiency of photosynthesis. In addition, one of the products of the oxygenation of Rubisco, 2-Phosphoglycolate (2-PG), is a strong inhibitor of several essential enzymes (SBPase and TPI) involved in the Calvin cycle.
Because the Calvin cycle is directly connected to the rate of carbon fixation, its regulation is crucial for overall metabolic stability, and is therefore imperative to the plant’s ability to cope with environmental fluctuations, e.g., changes in light intensity or temperature.
Research is being done to reduce photorespiratory losses, and therefore increase net photosynthesis, by genetic engineering.
In an article recently published in in silico Plants, Prof. Thomas Nägele and his PhD student Jakob Hernandez of Ludwig-Maximilians-Universität München use structural kinetic modeling to determine how necessary photorespiration elements are for the stability of the Calvin cycle.
Kinetic modeling describes the physical and biochemical processes in a biological system using equations, and its output provides information on the behavior of the system. Kinetic modelling relies on knowledge of parameter values which describe the affinity of an enzyme for its substrate, the speed of a reaction and the enzyme sensitivity to inhibition by regulatory molecules. The Calvin cycle is operated by 11 different enzymes that catalyze 13 reactions. According to Hernandez, “parameter values for enzymes in the Calvin cycle are scarce because they are difficult to measure and variable depending on factors like temperature and pH. We overcame this roadblock by combining kinetic modeling with structural modeling.”
Using standard kinetic modeling, the following parameter values are typically needed:
- maximum reaction rate,
- affinity an enzyme has for substrate, and
- affinity an enzyme has for inhibitor.
In structural kinetic modeling, alternative values are used that are easier to obtain:
- amount of substrates and products, and
- rates of enzymatic interconversion.
First, two structural kinetic model versions – including or excluding photorespiration – were run to determine if, and to what extent, photorespiration stabilizes the Calvin cycle. Stability indicates that essential metabolite concentrations are in a sustainable steady state allowing the cycle to continue. Hernandez and Nägele found that including photorespiration significantly stabilized the Calvin cycle and is therefore a necessary process in a changing environment.
Next, the authors investigated what elements of photorespiration were necessary for stability of the Calvin cycle. They first focused on the inhibitor 2-PG, which is created as a product of photorespiration. The model including photorespiration was run with varying levels of inhibition by 2-PG to determine its effect on Calvin cycle stability. Regulation by 2-PG was found to increase the stability of the Calvin cycle.
Hernandez concludes, “the value of this work lies in the estimation of a trade-off between carbon assimilation and metabolic stabilization under environmental changes.”
READ THE ARTICLE:
Jakob Sebastian Hernandez, Thomas Nägele, The trade-off function of photorespiration in a changing environment, in silico Plants, 2022; diac022, https://doi.org/10.1093/insilicoplants/diac022
This article is part of the special issue on Multiscale Modelling of Photosynthesis.





