In the last twenty years, advances in whole genome sequencing (the technique used to read the genetic code written in the DNA) have allowed researchers to shed new light on plant evolution and plant-specific biological processes. A recent perspective article highlighted that only 812 species out of half a million existing green plants have been sequenced. Nevertheless, the adoption of this modern technology has hugely increased our understanding of the origin and diversification of plant life or the molecular basis of important plant traits. For instance, the scientific journals Science and Nature recently published results of research projects that employed genomics to decipher the domestication of grapevine or to dissect the genetic control of seed characteristics in an important legume. For plants of agronomic interest, information on genomic sequences can be used in crop improvement programs to obtain cultivated varieties that are more resilient to stresses, require fewer inputs for growth, or have better nutritional value.
Cultivating Faba Bean, a Good Plant-based Alternative to Meat
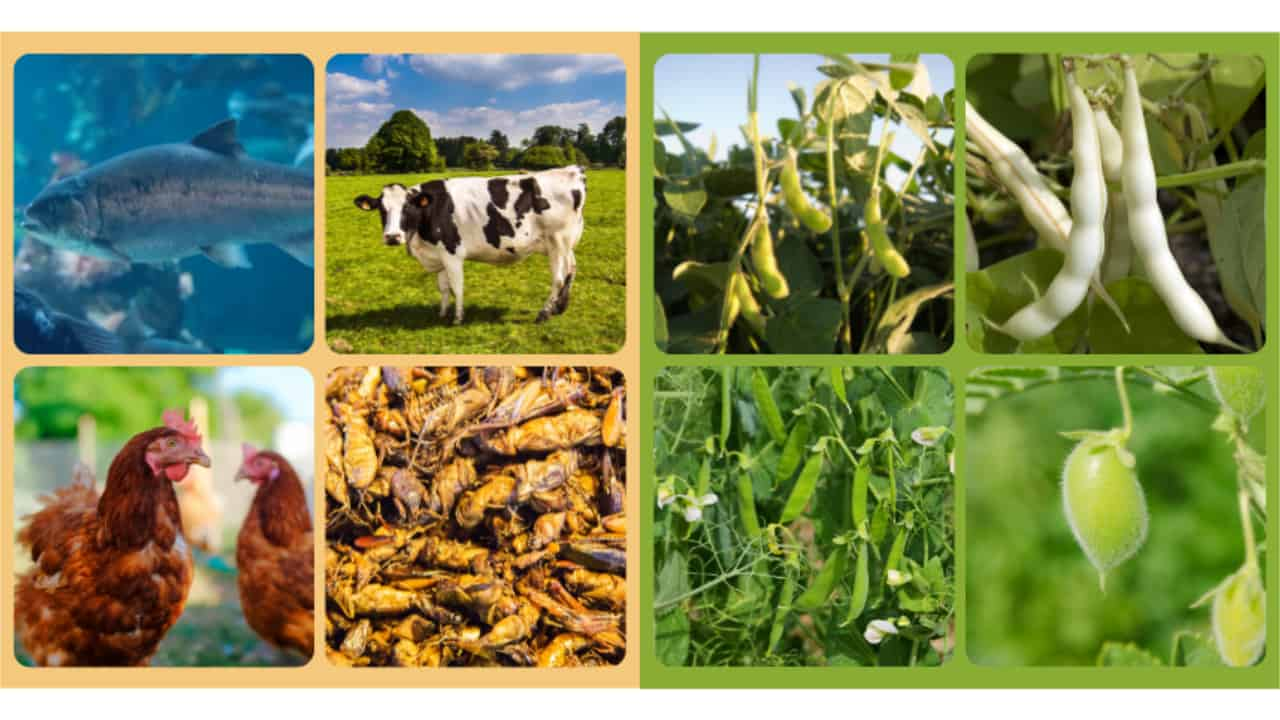
In a climate change scenario, producing plant protein as an alternative to animal protein could be a good strategy to reduce greenhouse gas emissions in agriculture (Figure 1). In particular, legumes – often referred to as “food with potential for significant health benefits” – can provide the cheapest source of high-quality vegetable protein. While Soybean (Glycine max) – the most important bean of the Fabaceae family – grows well under warm conditions, cool-season pulses (e.g., pea, lentil, chickpea) can be cultivated in temperate regions. Among pulses, Faba bean (Vicia faba) stands out due to its high adaptability, productivity, and nutritional value.

Despite the long tradition of faba bean cultivation in the old world, mysteries on its origin of domestication in the fertile crescent remain unsolved as no wild progenitor has been identified. In addition, a lack of genetic data and genetic material has hampered breeding schemes aimed at reducing the production of antinutrients (e.g., vicine, convicine) or balancing the content of essential amino acids.
Long Reads Revealed the Secrets of Faba Bean Giant Genome
Sequencing of the faba bean genome has long been hindered by its huge size: 13 billion nucleotides (13 Giga kb) distributed across 6 chromosomes. To have a better idea of this colossal dimension, the whole human genome is the same size as the biggest chromosome of faba bean!
Recently, an international team of researchers succeeded in sequencing the diploid genome (2 copies of 6 chromosomes) of faba bean by employing a new technology called PacBio HiFi long reads. The authors chose as reference an inbred line (Hedin/2) with favourable characteristics, such as high yield potential and early maturing habit. Interestingly, this line shows a high level of homozygosity (i.e., presence of two identical variants of a gene, one from each parent) thanks to a considerable degree of auto fertility (or self-pollination), despite being an allogamous species (i.e., reproduction by the cross-fertilization of pollen from another plant of the same species).
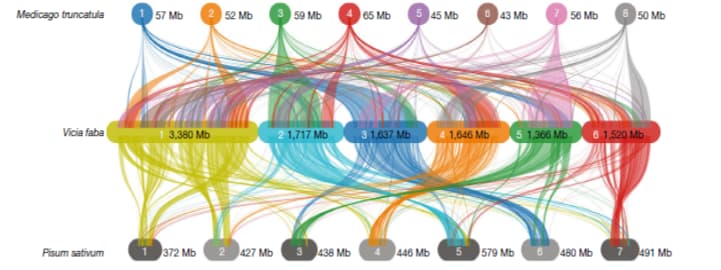
Jayakodi and coworkers discovered that the giant genome can be a consequence of multiple events – including the presence of huge intergenic regions, the expansion of repeated elements (such as transposons that account for 80% of genetic material), and tandem duplication of several genes. The authors also combined sequencing of genome and transcriptomes (all messenger RNAs transcribed from the corresponding genes) of 9 tissues to improve the annotation of 35000 protein-coding genes. As a curiosity, the distribution of genes across chromosomes is similar in Faba bean and phylogenetically close species Pisum sativum (Figure 2), despite the great difference in their genome size.
Agrigenomics for Breeders and Geneticists: A Wide Range of Applications
But what can molecular biologists and breeders do with genomic datasets? As proof of concept, the authors provide practical examples of application by carrying out Genome-wide association studies for important seed characteristics. Specifically, genomic information was used to set up genotyping assays to explore variation in a diversity panel of cultivated accessions and to identify which genetic variants cause the desirable traits. The faba bean genomic data can be further exploited for fast introgression of beneficial features such as reduced content of toxic compounds (e.g., alkaloid glycosides), increased protein bioavailability, or modulated accumulation of phytates in seeds.
Suggested Reading
Milestones in Genomic Sequencing (nature.com)
Representation and participation across 20 years of plant genome sequencing | Nature Plants
Decolonizing botanical genomics | Nature Plants
Green plant genomes: What we know in an era of rapidly expanding opportunities | PNASThe giant diploid faba genome unlocks variation in a global protein crop | Nature






