Crop modelling frameworks streamline modelling workflows from running simulations, to output visualization and model evaluation by providing handy tools that support the development, exchange, and reuse of crop model components. Modelers can use frameworks with little overhead for detailed comprehension of the underlying programming or need to access and modify the platform.
Frameworks give crop modelers the ability to:
- define models in a convenient way
- test alternative hypotheses in the same model
- exchange model components between different platforms
- organize complex simulations under various scenarios
- compare model output and performance
Postdoctoral researcher Kyungdahm Yun and Professor Soo-Hyung Kim, both at the University of Washington, introduce Cropbox, a declarative crop modeling framework, in a new article published in in silico Plants.
While many crop frameworks have been created, most recently, Crop2ML, CPlantBox, and yggdrasil, Cropbox is unique in that it provides a simple domain-specific language and useful tools for supporting an entire cycle of model development. “The framework was built to support an exploratory workflow with a minimal amount of coding in an interactive development environment. We hope it allows anyone interested in modeling to do “modeling” with little need to worry about technical details.”
Specifications
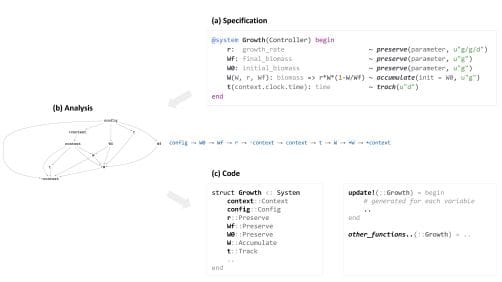
- It provides a declarative syntax resembling mathematical formulae for defining model components and variables (see figure).
- It automatically converts and validates units of measurement.
- It translates model specifications into Julia code. Julia is a new programming language that was designed for scientific computing and equipped with powerful packages that handle data management, unit conversion, statistics, and 3D visualization.
Simulations
- It provides a unified interface for running models and generating outputs in a standard format.
- It supports a programmatic construction of configurations for setting parameter values.
- It allows large scale simulations by using automatic parallel processing.
Evaluation
- It provides common metrics for model evaluation such as RMSE (root mean square error) and Nash-Sutcliffe model efficiency, a.k.a., model efficiency (EF).
- It supports automatic model calibration using a global optimization method.
Visualization
- It supports various plotting options out of the box for web-based Jupyter and text-based REPL environments.
- It allows interactive manipulation of model output via graphical user interface.
- It can support more powerful visualization (e.g., 3D rendering) via external Julia packages.
Kyungdahm Yun gives an overview of Cropbox at JuliaCon 2022.
The authors provide three example applications where previously existing models written in C++ were reimplemented in Cropbox: a coupled C3 and C4 leaf gas-exchange model, a process-based crop model for garlic, and a 3D root architecture model. These examples demonstrate streamlined workflows and less code using Cropbox.
Yun concludes “We anticipate that Cropbox will become a versatile toolbox for crop modelers to build, improve, and apply crop models in ways that have yet to be seen. We also hope that Cropbox will become an agent that blurs the lines between model users and model developers, with fewer technical hurdles.”
READ THE ARTICLE:
Kyungdahm Yun, Soo-Hyung Kim, Cropbox: a declarative crop modeling framework, in silico Plants, Volume 5, Issue 1, 2023, https://doi.org/10.1093/insilicoplants/diac021
The open-source framework, Cropbox, is freely available at https://github.com/cropbox/Cropbox.jl. There, readers can also find additional examples of use.




