Anyone who experimented with sourdough starters during the Covid-19 pandemic knows that bread bakers prize wheat flour with high levels of protein. Proteins ensure that the bread dough is elastic and extensible. These characteristics allow dough to better trap carbon dioxide, causing it to rise more vertically and making the baked loaf light and airy.
The importance of wheat extends beyond hobby bakers. Wheat is a staple food that provides around 20% of all calories and protein consumed by humans. Demand for wheat is expected to increase by 60% by 2050 while declines in yield attributable to diminishing water resources and heat due to climate change are already being seen.
Scientists are scrambling to develop a wheat that can produce high yields despite the climate crisis. However, it is difficult to select for complex traits like yield without also affecting seemingly unrelated traits – an increase in wheat yield is negatively correlated with protein content (which is essential for human nutrition and also important for a good sourdough).
Genomic selection models can illustrate how wheat yield can be increased without reducing protein content.
National Institute of Agricultural Botany (NIAB) graduate student Nick Fradgley and colleagues used genetic models to determine how to optimize the selection of multiple traits in breeding in a new paper published in in silico Plants.
The study used a newly developed mapping population, called MAGIC, of over 500 inbred lines descended from sixteen historical bread wheat varieties that have been extensively genotyped and phenotyped. The use of this population and associated data allowed the researchers to investigate the genetic basis for phenotypic variation.
The authors first investigated the complex relationships among traits. This was accomplished using software that examined how 72 traits correlated with each other for the 500 lines and two years of phenotypic data. As expected, they found a strong negative correlation between protein content and yield.
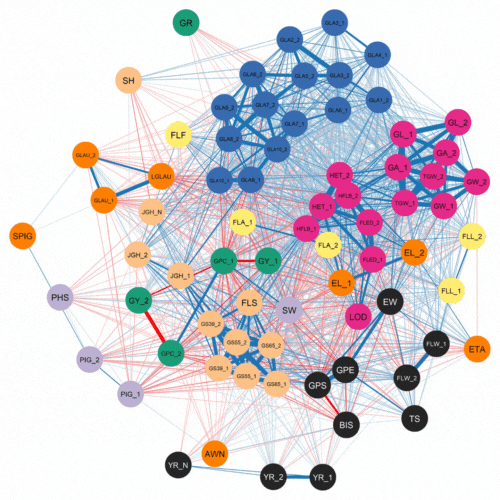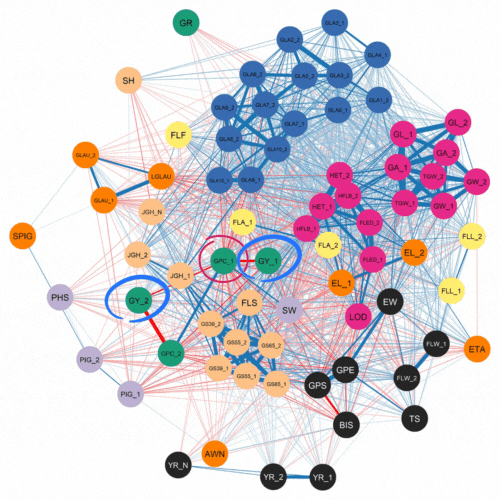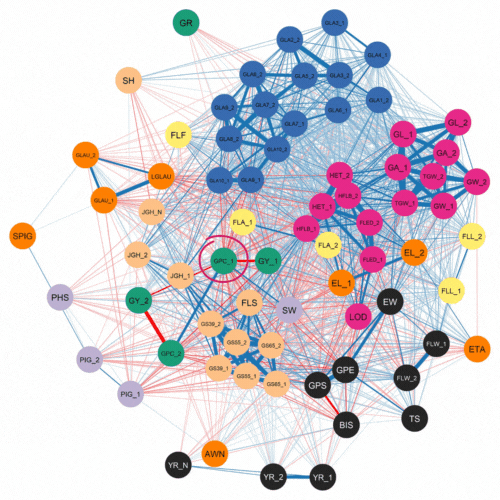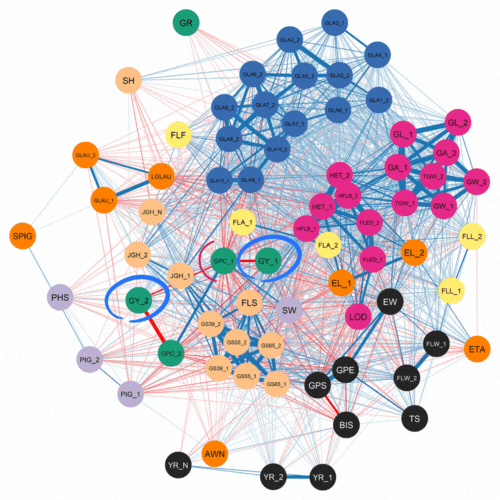
Numerous genomic selection studies have analyzed traits separately using single-trait analyses. This technique treats each trait as unrelated. In contrast, multi-trait models can incorporate information about correlated traits which are controlled by a common genetic effect. This phenomenon, where one gene can influence two or more seemingly unrelated phenotypic traits is known as pleiotropy.
To highlight the importance of pleiotropic effects in prediction accuracy of complex traits like grain yield, the authors compared the accuracy of single-trait and multi-trait genomic prediction models.
Data from 90% of the lines was used to train and optimize the models. The remaining 10% was used as a test dataset. The models were asked to predict the value of multiple related traits using only genetic data from the test dataset. The authors then compared the predicted with the measured trait values from the test dataset. They found that compared to single-trait models, multi-trait models had higher prediction accuracy for almost 90% of traits, improving grain yield prediction accuracy by 3-52%.
Next, they investigated the potential for achieving long-term genetic gain in grain yield by simulating a long-term breeding program using two methods. For this, they used the breeding strategy recurrent genomic selection, which is repeated cycles of selection and breeding over multiple generations aimed at gradual genetic improvement of one or more traits.
They simulated long-term breeding with two separate selection goals:
- improvement of a single trait – grain yield, or
- improvement of multiple traits simultaneously – grain yield and other traits of interest.
For each of the 20 breeding cycles, the model selected lines for crossing based on their advantageous phenotypes, which were predicted from their genotype. Traditional wheat breeding program cycles generally last five years. Therefore, these simulations of 20 cycles of recurrent selection represent an equivalent of over 100 years of traditional wheat breeding.
When the goal was selection based on grain yield, they found rapid genetic gain in yield with, as expected, a corresponding rapid decrease in grain protein content.
However, with the use of multiple trait selection genetic gain in both yield and protein content was possible to some extent. The rate of genetic gain in grain yield was slowed but the selection succeeded in increasing both desirable traits, indicating that it is possible to optimize antagonistic traits through appropriate selection.
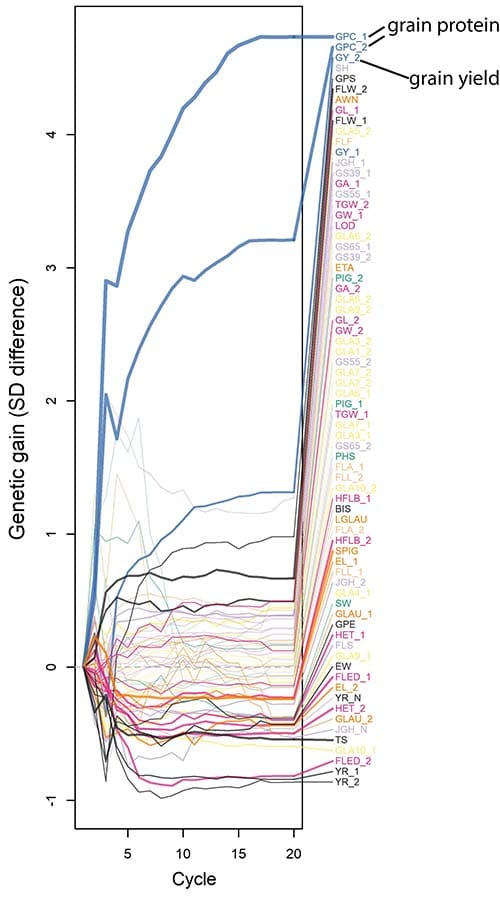
These results show that there is promise in meeting high protein wheat yield goals with collaboration between modelers and wheat breeders.
The authors conclude, “further genetic gain in current breeding programmes will therefore likely be achieved through optimisation of small and complex genetic effects. These findings highlight the important role of models to maximize further genetic gain in the future.”
READ THE ARTICLE:
Nick Fradgley, Keith A Gardner, Alison R Bentley, Phil Howell, Ian J Mackay, Michael F Scott, Richard Mott, James Cockram, Multi-trait ensemble genomic prediction and simulations of recurrent selection highlight importance of complex trait genetic architecture for long-term genetic gains in wheat, in silico Plants, Volume 5, Issue 1, 2023, diad002, https://doi.org/10.1093/insilicoplants/diad002






