In the last decades, ecologists have increasingly used a trait-based approach, which means they are using morphological, physiological or phenological characteristics of organisms to interpret and understand ecological dynamics. For seed ecologists, such traits usually come in the form of seed size measurements and descriptions of seed shapes and appendices. Obtaining such morphological characteristics can be a highly time-consuming process that involves long days in the microscope and deciphering the best way to classify seeds into subjective shape and colour classifications. Image analysis is the most natural solution for this task, as computers could be programmed to extract information from seed pictures. Still, the most common approaches, such as thresholding and deep learning, either struggle with the high diversity of seed morphologies or require sophisticated computational resources.
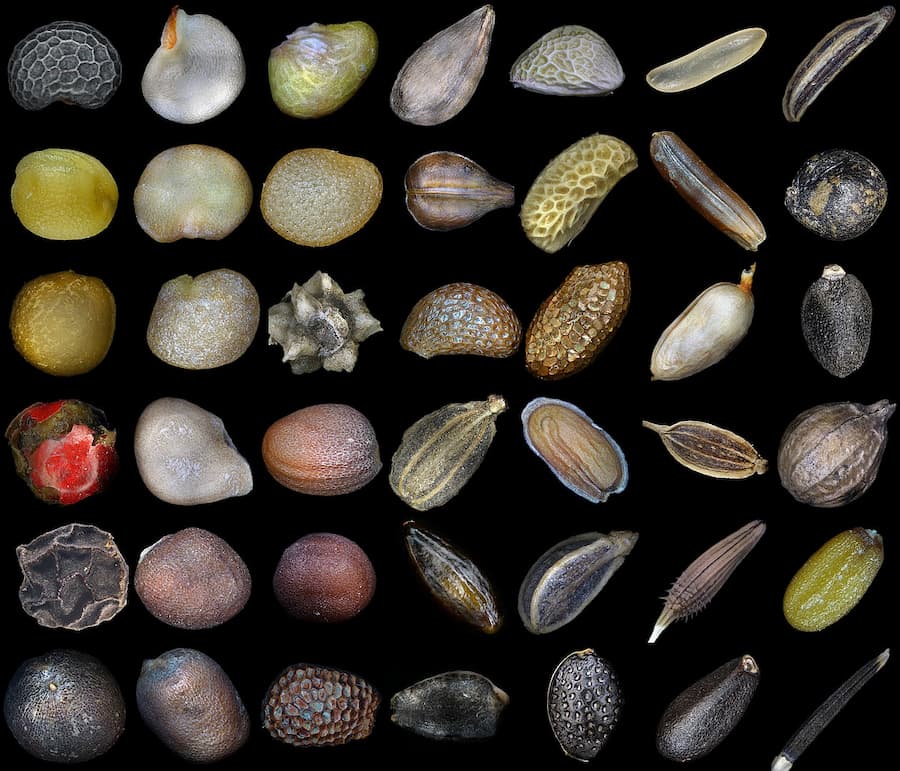

In response to these challenges, a research team led by Dr Roberta L. C. Dayrell (@RobertaDayrell on Twitter) created Traitor –an open-source software that automates the extraction of seed size, shape, and colour traits from images. Unlike other available tools, Traitor employs an unsupervised segmentation technique that enables the automated separation of seeds from high-contrast backgrounds without training or manual adjustments. Moreover, the software is designed for processing hundreds of images from varied taxa, enabling large-scale analysis that was previously unfeasible.
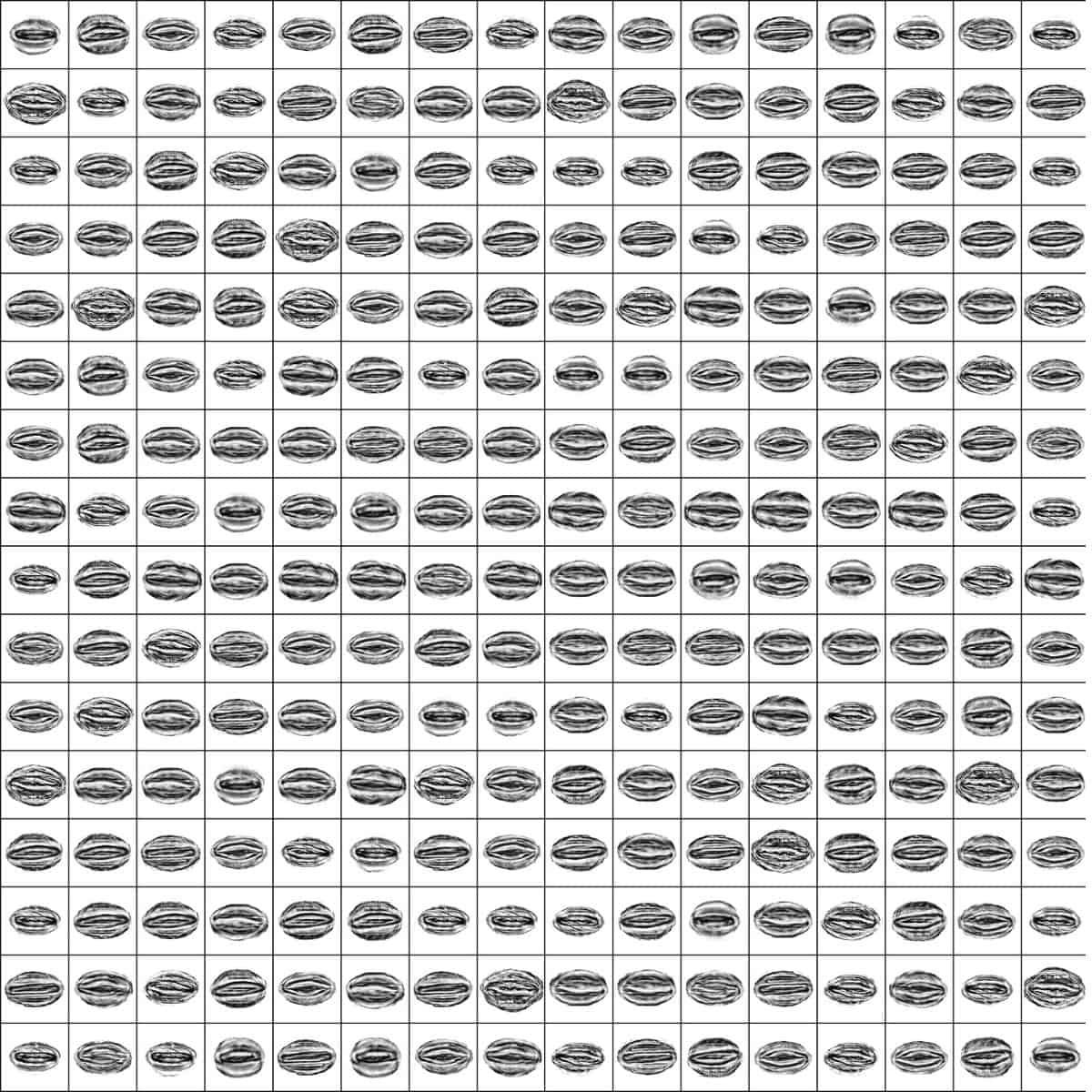
To use Traitor, the user only has to follow four steps. First, seeds must be separated from debris and other appendices, such as hairy structures and awns, that could complicate image acquisition and processing. Then, seed images are taken using a scanner with a homogeneous colour background that contrasts with the seed ones. Such scans should include a colour chart containing standards for the next step, where the images are processed for colour standardisation, and seeds are automatically separated from their background. At the end of the processing phase, a silhouette is created for each seed, which is then aligned to be in relatively the same position as the others. Finally, the contours are used to extract all the traits. One could think that all these processes would require lots of commands, but the truth is that Traitor can do all of this using only three commands!
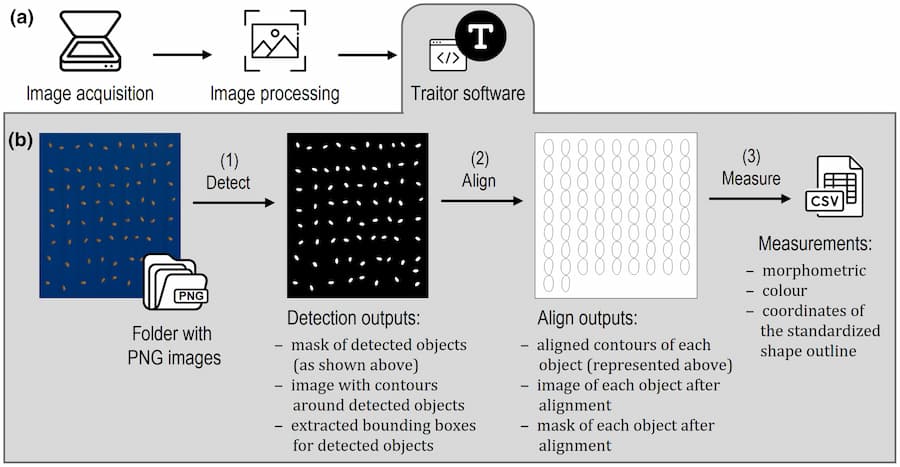
In an interview with Botany One, Dayrell commented that the main challenge encountered during the development of Traitor was defining the specific traits to be extracted. In her own words, the team’s goal “was to extract traits that could effectively communicate interpretable information about the morphological characteristics of seeds.” She continues, “This proved to be a complex task, particularly when dealing with shape and colour, which are intricate and multifaceted features.” In the end, this first version of the software includes twelve traits, including morphometric (such as length, area and perimeter), colour and shape measurements.
To verify the accuracy of the measurements of their program, Dayrell and her team used DiasMorph –a trait and image database of seeds from Central Europe with over 1200 taxa– and compared the measurements obtained by Traitor with ones obtained manually. The validation results leave no doubt about the accuracy of Traitor: the manual and automatic measurements coincided by about 98%! However, Traitor does not only work to extract this kind of morphological information but has excellent potential for obtaining data such as colour description and shape classification.
Seed colour is valuable, for example, to distinguish different species and has been associated with processes of great ecological importance, such as seed persistence in the soil. Yet, given the infinite number of colours that exist, it is always tricky to compare colours between observers. One might think you can solve this problem by classifying colours into broad categories –such as red, brown and grey seeds– but that would mean losing subtle variations. Traitor overcomes these challenges by extracting the sRGB values matching the seed’s colours. These values, which are universal and unique for each colour, can then be used in quantitative analysis that can span from describing colour variation within clades to the evolution of seed colour, paving the way for new and exciting research avenues and applications.
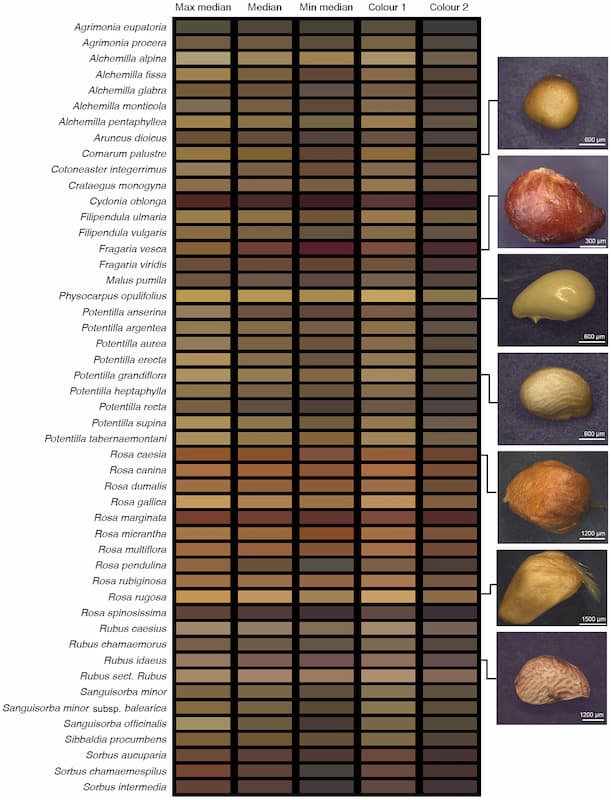
Similar challenges appear when researchers want to describe shapes, as the limits between one and another can be subjective and abstract. For example, when does a seed stop being “round” and start being “oval”? What is the boundary between “oval” and “ovoid” seeds? Well, Traitor frees us from such concerns, as it can produce quantitative values on seed shapes, including circularity and height-to-width ratio, allowing us to analyse seed shapes objectively. For instance, Dayrell and her team used these data to characterise the seeds of Carex species found in DiasMorph and classified them into eight different shape categories based on aspect ratio and symmetry. Notably, some species were consistently assigned to a given shape class, illustrating their approach’s potential for building seed identification tools.
Considering all these outstanding functions, Traitor promises to be a handy tool for advancing seed ecology, as it will allow the automated extraction of various seed traits in a user-friendly and time-efficient manner. Moreover, the quantitative descriptors of shape and colour will enhance a more accurate and interpretable description of such characteristics. Its successful validation with the DiasMorph database proves its high potential of producing large, reliable datasets of seed traits, a resource that is missing in most parts of the world, especially the Tropics. As Dayrell told us in the interview, “It is worth noting that this method of trait extraction is cost-effective, requiring only a computer, a scanner, and a high-contrast canvas. This affordability makes it particularly valuable for research efforts in labs where resources may be limited.”
These results are important outside ecology as well. Dayrell pointed out that, for example, this software has great potential for plant breeding. She argues that “identifying and selecting seeds with desired traits play a crucial role in plant breeding programs. Traitor‘s output provides valuable information for evaluating and sorting seeds based on their morphological attributes. This capability enhances the efficiency and effectiveness of plant breeding efforts, facilitating the identification and propagation of seeds with desired characteristics”. She also highlighted the potential role of Traitor in generating data that can fuel the development of intuitive and user-friendly identification tools that can be used in various disciplines, such as ecological restoration, archaeobotany, and ethnobotany. One thing is sure: the possibilities are endless, and Traitor seems up to the challenge. If you ask me, I can’t wait to give it a try!
READ THE PAPER
Dayrell, R.L.C., Ott, T., Horrocks, T., & Poschlod, P. Automated extraction of seed morphological traits from images. Methods in Ecology and Evolution. Available at: https://doi.org/10.1111/2041-210X.14127





