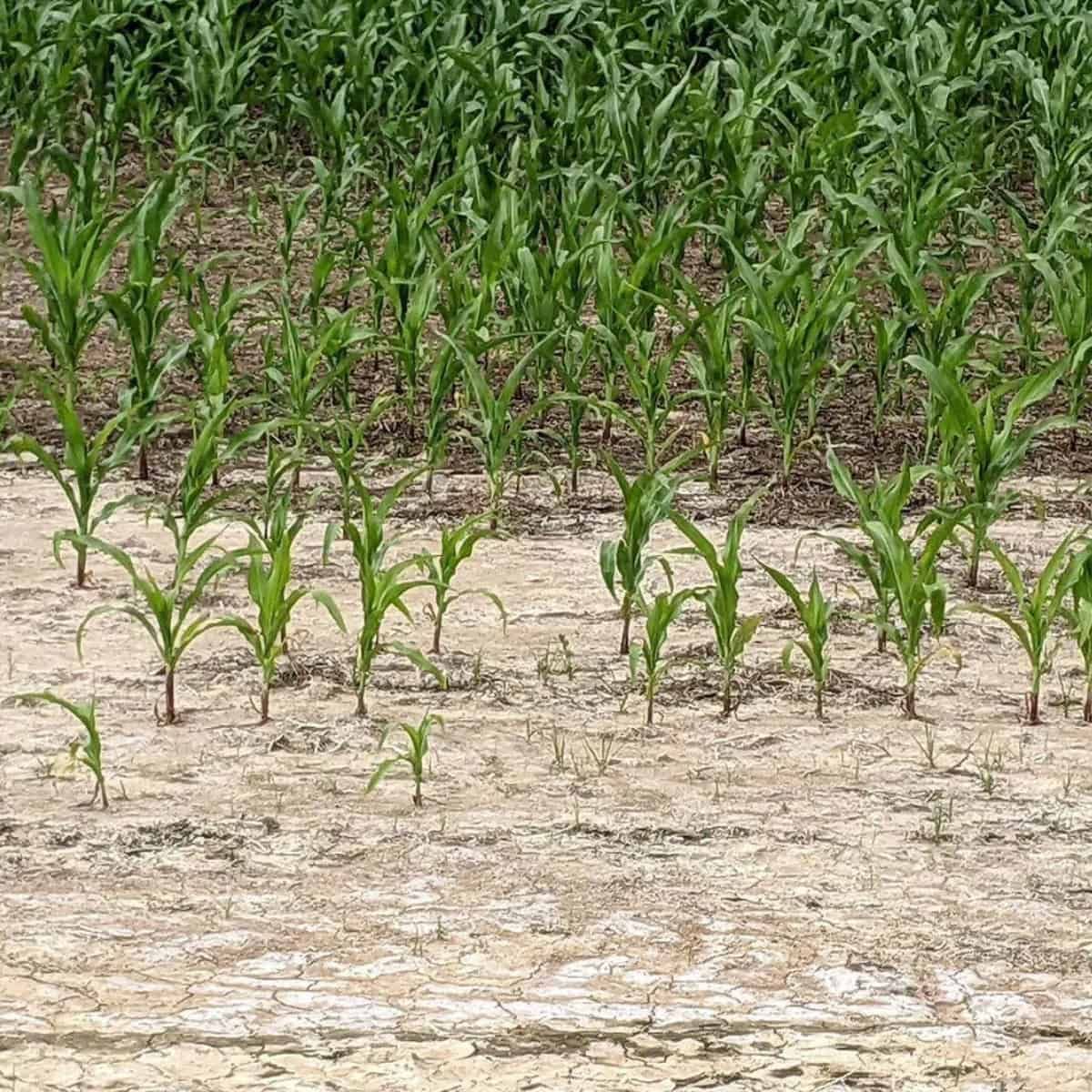
In the quest to better understand plant life, Richard Harwood’s recent work, published in AoB PLANTS, examines the concept of a ‘virtual leaf’. He leverages cutting-edge microscopy techniques and computational modelling to study leaf physiology in three dimensions (3D), a leap from traditional 2D approaches.
The concept of a ‘virtual leaf’ aims to replicate the complex physiology of a leaf in a digital environment, allowing for computational simulations of processes like water evaporation and carbon dioxide (CO2) transport. These simulations provide insights into how the unique 3D structures of different leaf tissues influence carbon and water exchanges, ultimately affecting processes like photosynthesis and respiration.
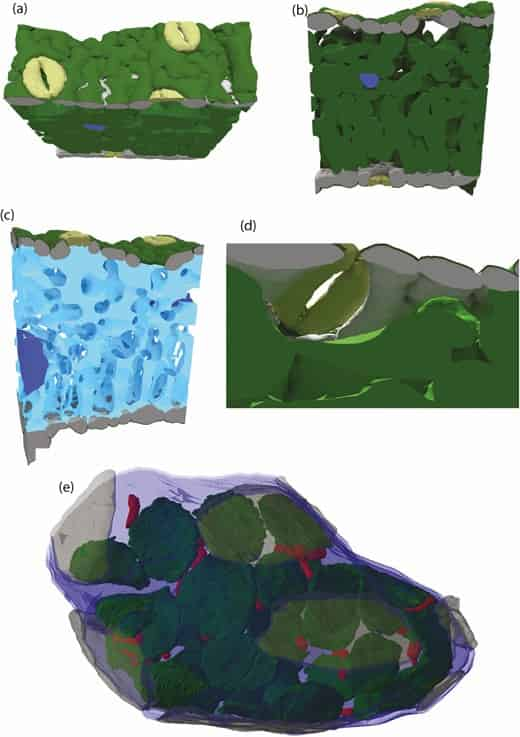
Harwood also investigates mesophyll conductance in 3D, which involves the passage of CO2 from the tiny air spaces in a leaf to the photosynthetic cells. His research points out that the size, shape, and arrangement of cells within the leaf significantly affect this process. Previous approaches assumed a uniform distribution of carbon dioxide within the leaf, failing to account for the leaf structure’s real-life variability and complex biophysical properties.
A ‘virtual leaf’ could help researchers visualize water transport systems within leaves and even track the movement of carbon and water inside a leaf in real-time. For instance, Cicer arietinum or chickpea leaf’s complex structures and responses to light could be captured with this technology, creating a more detailed understanding of plant response to environmental changes.
Importantly, Harwood emphasizes that a ‘virtual leaf’ should also account for dynamic, temporal changes like responses to drought. He commends the work of Xiao and his team for their ‘eLeaf’ model, which he sees as a valuable step towards creating a comprehensive ‘virtual leaf’. He writes:
It is important to keep a grounded perspective when attempting to capture in a laptop physiological process that evolved over millennia. As such, research efforts that contribute to the advancement of a ‘virtual leaf’ need to be assessed in terms of how they improve our limited understanding of plant physiology, not how they fail to capture complex reality. Xiao et al. (2022) exemplify an excellent approach to creating a ‘virtual leaf’. The authors use different microscopy modalities (micro-CT, confocal and transmission electron microscopy) to capture leaf anatomy at different spatial scales. This anatomical information is then pooled together and used to create a 3D structure that is representative of the microscopy. Whilst still somewhat ‘cartoon’ like in nature it is an excellent concession between 3D meshes that can be used in modelling software and authentic 3D anatomy. The authors validated their modelling assumptions with experimental data and termed the final product ‘eLeaf’.
Xiao et al. (2022) utilized ‘eLeaf’ to discover that changing leaf porosity had little influence on photosynthetic performance which aligns with the observations from Theroux-Rancourt et al. (2021) that cell size is much more important than porosity. It is worth noting that Theroux-Rancourt et al. (2021) took 3D anatomical data to estimate 2D CO2 diffusion and Xiao et al. (2022) took 2D anatomical data to estimate 3D diffusion. As 3D modelling approaches evolve, we will gain a better understanding of what level of detail is needed to answer specific research questions. Advancements in imaging techniques and artificial intelligence as a tool for segmentation (Théroux-Rancourt et al. 2020; Heinrich et al. 2021) will result in more refined and anatomically accurate 3D models. As these 3D models unravel the complexities of leaf physiology a ‘virtual leaf’ will emerge, representing a powerful tool that merges microscopy, modelling and experimental data.
Harwood 2023.
READ THE ARTICLE
Harwood, R. (2023) “Creating a virtual leaf,” AoB PLANTS, 15(3), p. lad033. Available at: https://doi.org/10.1093/aobpla/plad033.




