There are sunflower festivals around the world. You can wander among fields of radiant flower heads but, even with your eyes wide open, you can miss the signals the sunflowers are sending. Flowers are tools that plants use to attract pollinators. If pollinators were urban hipsters, plants would have flowers shaped like coffee shops, with an attractive display guiding them to the door. For sunflowers, pollinators are insects, and so they display something eye-catching for them. But insects’ eyes see the world differently.
The key difference is, unlike humans, they can see ultraviolet light. Sunflowers take advantage of this by producing UV-absorbing pigments in their flower heads. What you see as a ring of gold around a dark centre is, for insects, a much more complex series of rings. There’s the outside floret, which is sterile and only works for display to insects. Then there’s the internal florets that actually get pollinated. That, and the UV patterning makes the flower head more an archery target or bullseye. It might be invisible to us, but understanding this patterning is important, for pollination and seed production. New research by Brook Moyers and colleagues has found the secret to this pattern in the plant’s DNA.
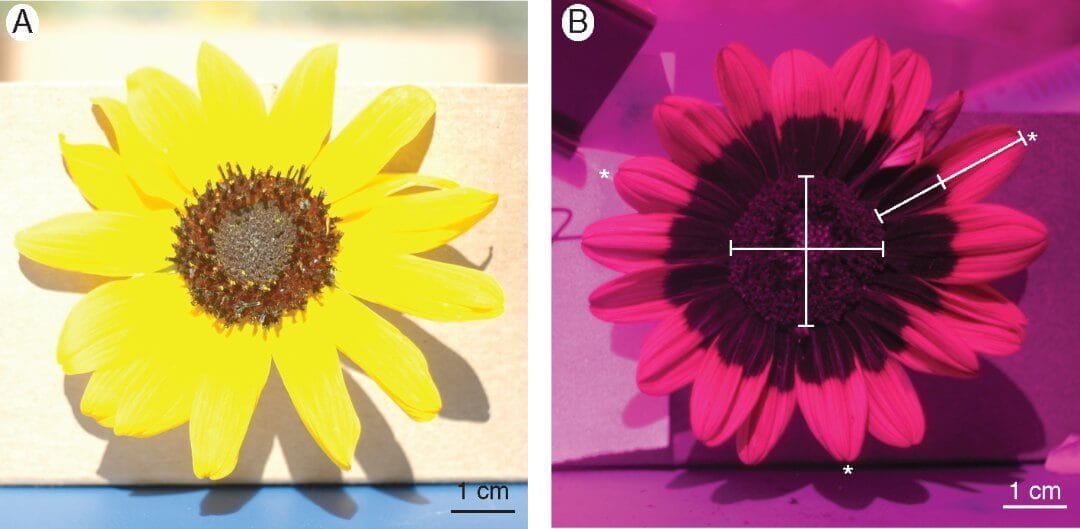

Moyers et al. examined quantitative trait loci: these are sections of DNA in the genome that have correlations with variations in the phenotype, the physical form of the plant. What they looked for were single nucleotide polymorphisms, usually called SNPs (pronounced ‘snips’). Nucleotides are the molecules that are the building blocks of the DNA, and they’re sometimes thought of as the ‘letters of DNA’ that make up the genome. SNPs are where these letters differ between organisms of the same species. For example you might have a sequence like this:
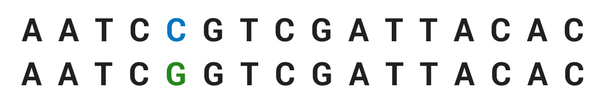
That difference is a SNP. They happen around every 300 molecules, and can mark genetic variation.
Working with Helianthus argophyllus, a sister species of the domestic sunflower, Moyers et al. found six quantitative trait loci based on SNPs that affect the display of the sunflower in the ultraviolet spectrum. They don’t identify specific genes, but with the QTL approach, they don’t have to. They also found that the quantitative trait loci they were studying for bullseye and flower head size were in the part of the genome that defined flowering time. This means that somewhere in this section of the genome there is likely to be a gene that affects multiple factors in the flower head.
Another feature they found is that the part of the genome that makes the ultraviolet pigment is unrelated to the part that defines the size of the ultraviolet bullseye. This matters because the size of the bullseye might be connected to the size of the flower head, but the intensity of the pattern is not. If you’re cross-breeding plants to get a bigger and better display for pollinators, that means there’ll be two different genes you have to account for.
Embed from Getty Imageswindow.gie=window.gie||function(c){(gie.q=gie.q||[]).push(c)};gie(function(){gie.widgets.load({id:'Ak8Bx5SPQfxeTFRaCW0Y0w',sig:'j2EMYYhQ1xbXyLoiAnv5V2TFKPN03e5WZa2JAMfmLp0=',w:'594px',h:'395px',items:'94824737',caption: true ,tld:'co.uk',is360: false })});</div> <p>Sunflowers aren’t just pretty, they’re an economically important crop, so there’s a demand for better and more productive varieties. Knowing how these quantitative trait loci interact with the floral display means you can identify better varieties of sunflower for cross-breeding. Given the number of <a href="https://en.wikipedia.org/wiki/Sunflower_oil#Uses">uses for sunflower oil</a>, a better supply based on this research might well make a lot of products less demanding on your bank account.</p> </body>




