Cotton fiber is the most produced non-food crop in the world according to the FAO. Its production contributes significantly to the economies of many developing countries and to the livelihoods of millions of rural smallholders worldwide.

Modern cotton varieties were derived from domestication and improvement processes that trace back thousands of years. The major change over this time was that cotton fibers have become longer, finer, and stronger.
Modern efforts to engineer cotton to further improve fiber quality are hampered by insufficient knowledge about the underlying mechanisms controlling fiber characteristics.
Dr. Daniel Szymanski, Professor of Botany and Plant Pathology at Purdue University and Dr. Joe Turner at University of Nebraska developed realistic simulations which allowed them to analyze the cellular components that control cotton fiber diameter and length.
Each cotton fiber is an individual elongated cell that grows on the surface of the developing seed coat. This makes them easy to purify and image using advanced microscopy techniques, making them a powerful single cell model for insight into plant growth and development, cell wall assembly, and cell morphogenesis.
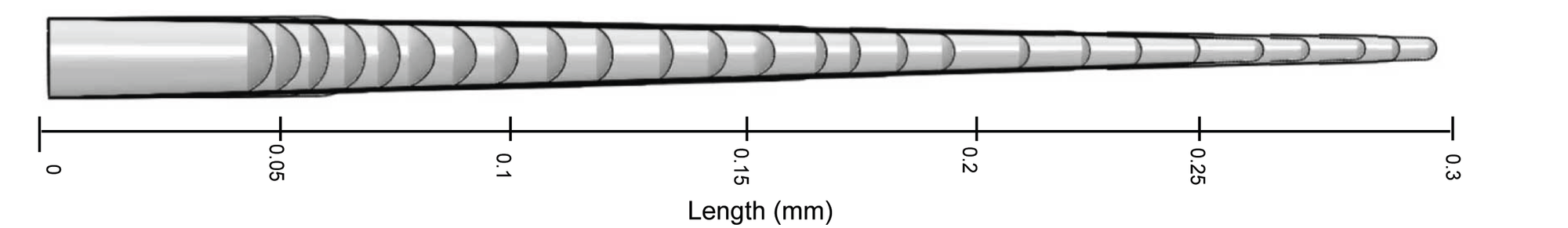
The anatomy of the fibers determines final product quality. Dr Szymanski explains, “the diameter of the fiber contributes to its fineness in terms of a higher thread count and a soft silky feel. At the start of cell development, the fiber rapidly tapers, and the resulting diameter appears to be maintained as the fiber elongates. We therefore focused our efforts on the mechanisms of cell wall tapering because the extent and duration of tapering determines its diameter.”
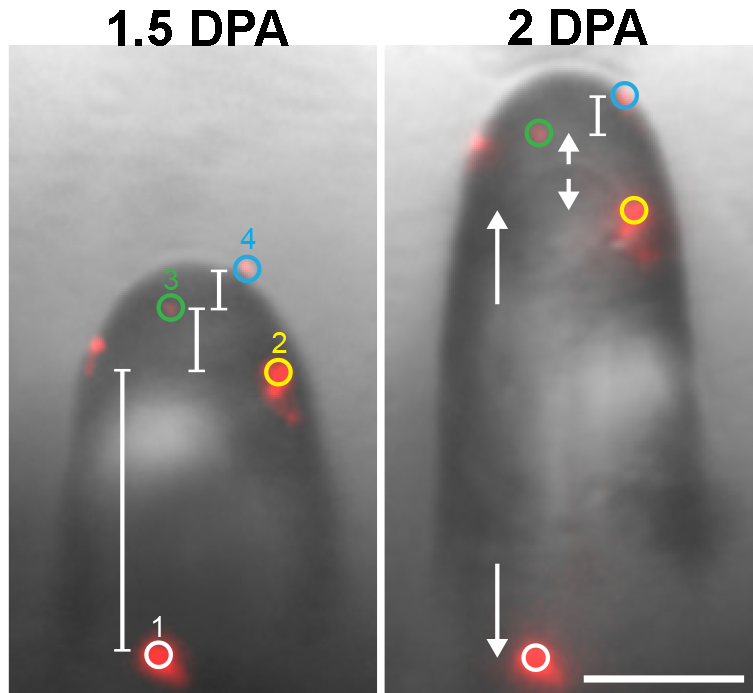
Live-cell microscopy was used to determine the precise developmental timeline and extent of fiber tapering measured in days after anthesis. This included measurements of local cell expansion rates and radius of curvature of the fiber tip (a proxy for fiber diameter) during early fiber development.
Their investigation focused on the cytoskeleton structures because they enable cells to generate and maintain highly polarized (asymmetrical) cell shapes. Cytoskeletons can be made of actin or microtubule filaments; both cooperate to mediate tapering in cotton fiber. To analyze actin networks in tapering cells, they used a combination of live cell imaging and particle tracking. Through localization of the actin roadways in the cell, the authors determined how the cell distributes the delivery of raw materials throughout the expanding cell surface.
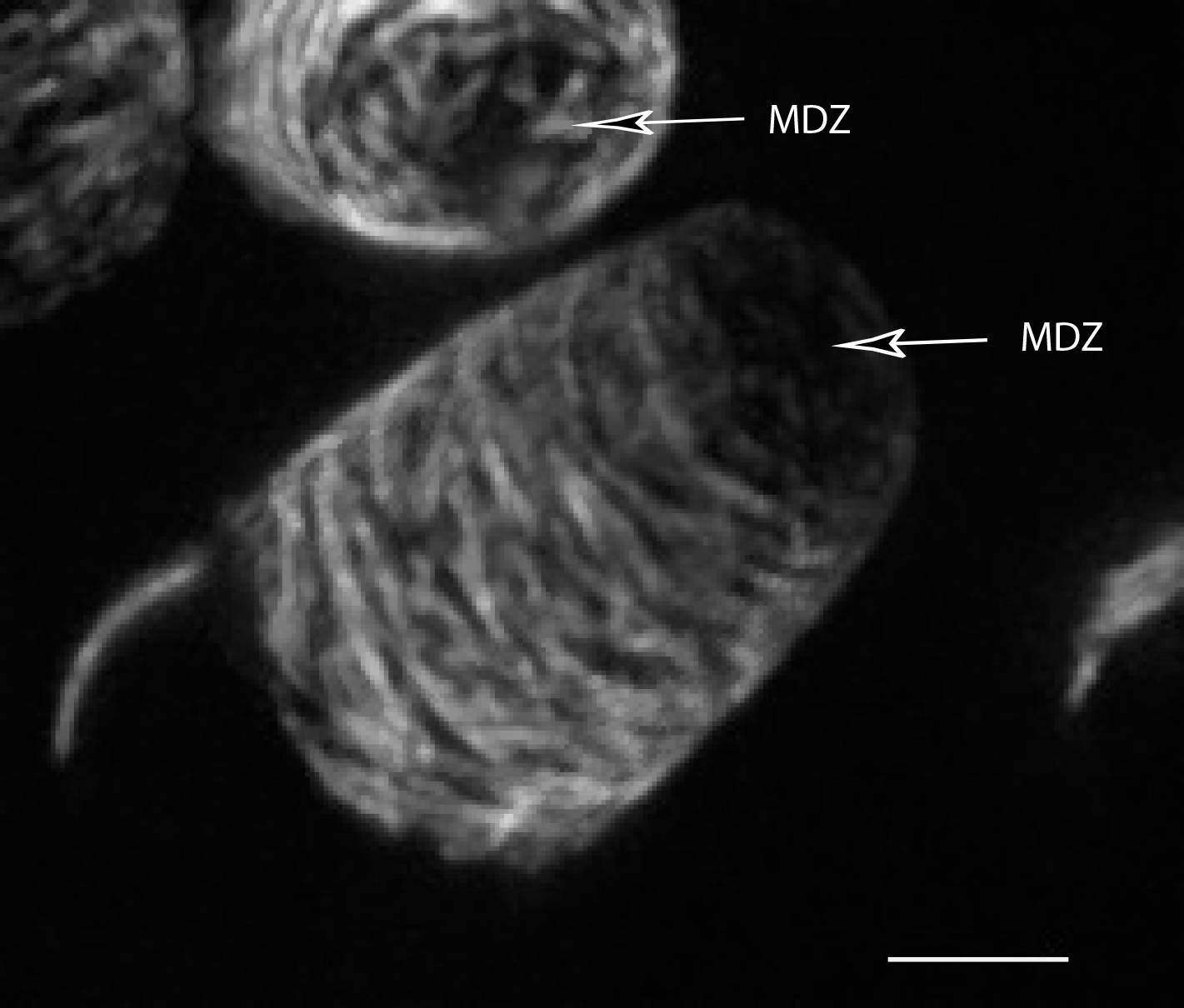
Interestingly, they found that the microtubule cytoskeleton, which patterns the orientations of cellulose microfibrils in the growing cell wall, includes an apical microtubule depletion zone and highly organized transverse microtubules.
The authors created a finite element model of cotton fiber growth to determine what material properties in the wall are required to enable fiber tapering and fiber elongation without cell swelling. These simulations revealed a strict requirement of transverse fibers to restrict radial swelling and an isotropic zone of randomly oriented fibers in the apex to cause the cell diameter at the apex to become progressively restricted during growth. The model also provided a reliable way to predict the magnitudes and directions of tensile forces in the wall. These simulations indicate that force magnitude and orientation feed back on the microtubule system to allow the cell to integrate cell shape with microtubule-dependent cell wall patterning. Central to the creation of the model were the multi-scale experimental data described above.
According to Szymanski, “the combination of finite element model simulation and modern cell biology provides a knowledge base to enable fiber engineering. The key challenge now is to identify the genes and modules that underlie important fiber traits like diameter, twist, and length”.
READ THE ARTICLE:
M Yanagisawa, S Keynia, S Belteton, J A Turner, D B Szymanski, A conserved cellular mechanism for cotton fiber diameter and length control, in silico Plants, 2022;, diac004, https://doi.org/10.1093/insilicoplants/diac004






